Mice and Housing Conditions
All experiments involving live mice were carried out under protocols sanctioned by the IACUC at both Brigham and Women’s Hospital and Stanford University. Mice lived in standard conditions with unlimited access to food and water, on a light-dark cycle of 12 hours each.
For brain tumor allograft experiments, NSG mice (NOD-SCID-IL2R gamma chain-deficient, from the Jackson Laboratory) were the subjects used, with male and female mice utilized equally. Following IACUC guidelines, signs of morbidity rather than the size of the tumors were the criteria for ending experiments. Mice would be euthanized when signs of neurological issues appeared or if they lost more than 15% of their body weight. For optogenetic stimulation studies of the premotor circuit (M2), Thy1-ChR2; NSG or WT; NSG mice were employed.
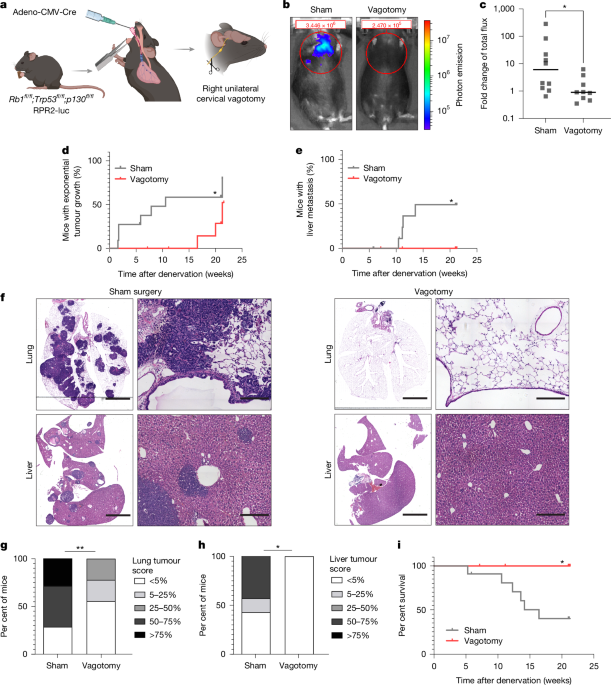
In lung tumor experiments, euthanasia occurred upon noticeable symptoms of illness, like difficulty breathing or loss of weight beyond 15%. No operational limits were exceeded during these trials. Genetic mouse models such as Rb1fl/fl;Trp53fl/fl;p130fl/fl expressing luciferase (RPR2-luc) were used as previously detailed. These mice naturally developed lung tumors and further metastases post-administration of Adeno-CMV-Cre (from University of Iowa Vector Core) at two months old.
Intracranial Allografts
All small cell lung cancer (SCLC) brain allografts were conducted as previously described. Essentially, a single-cell suspension was derived from cells cultured from either 16T-mGFP or NCI-H446-GFP SCLC neurospheres immediately prior to surgery. Mice aged 21-35 days were anesthetized and placed in a stereotactic frame. The skull was exposed through a midline incision under sterile conditions, and 70,000 cells in a 3 µl sterile solution were injected into the M2 cortex using a digital pump.
In Vivo Optogenetic Manipulation
For optogenetic stimulation of the M2 region in Thy1-ChR2; NSG or WT; NSG mice, a singular stimulation protocol was followed as outlined. A fiber optic connection was established either concurrently with or a week after the SCLC allografts. After a recovery period of one to two weeks, mice were connected to a laser system for light pulses for 30 seconds every minute over a session of 30 minutes. Mice were euthanized for further histological analysis 24 hours after stimulation.
Immunohistochemistry of Patient Tissue
Patient tissue samples were handled according to institutional review board-approved protocols. Immunohistochemistry was performed on fixed, paraffin-embedded sections following standard procedures. Various antibodies were utilized, such as mouse anti-Ki67 and anti-neurofilament. Ki67 staining was carried out using an automated stainer, while neurofilament staining was performed using a different automated system. The proliferation index was determined by counting the proportion of Ki67-positive cells relative to the total cell number in a designated area.
Single-Cell Sequencing from SCLC-Neuron Co-Culture
The 16T SCLC cells were either cultured solo or alongside neuronal co-cultures, with collection occurring in specific buffer conditions. GFP-negative cells were also collected as controls. Live-cell labeling was utilized and sorted cells were prepared for sequencing. The protocol followed standard methods for producing libraries suitable for high-throughput analyses.
Data analysis was performed using software designed for single-cell sequencing with filters applied to ensure quality control. The analysis involved principal component analysis (PCA) and clustering, allowing the identification of differentially expressed genes.
Sample Preparation and Image Acquisition for Electron Microscopy
For electron microscopy, NSG mice received engraftment with either 16T-mGFP or NCI-H446-GFP cells. Following three weeks, mice were sacrificed and subjected to transcardial perfusion using a fixative solution. Post-fixation procedures included dehydration and embedding, preparing samples for sectioning and imaging. Immunocytochemistry was also conducted to identify specific proteins within cells.
Statistical Analyses
Statistical evaluations were performed with GraphPad software, confirming Gaussian distribution where applicable. Two-tailed tests were utilized for parametric data, while non-parametric tests were applied as needed. Kaplan-Meier survival analysis was used to analyze survival data, with various graphical representations elaborating on results.