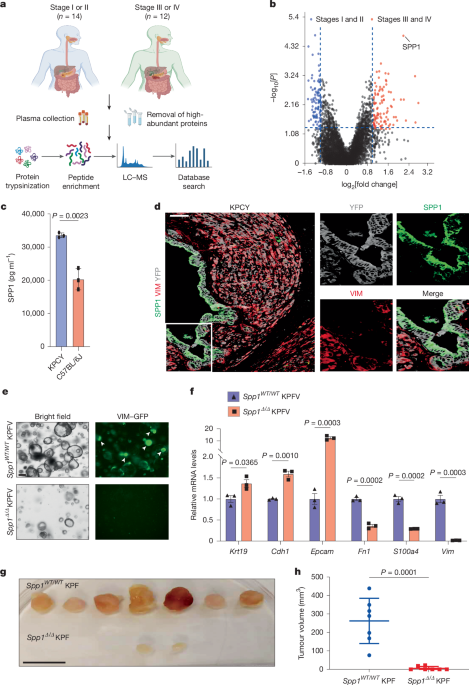
Blood samples from patients with pancreatic cancer
Blood samples from 26 patients diagnosed with pancreatic cancer were taken at the Seventh Affiliated Hospital of Sun Yat-sen University. Plasma was then extracted from the samples for proteomic analysis. All participants provided informed consent, and the research project received ethical approval from the hospital’s ethics board. Clinical data, including age, sex, TNM stage, and histology, were extracted from the participants’ medical records.
Generation of patient-derived organoids
Human pancreatic tissues were sourced from surgery patients at the same hospital. Consent was obtained from all individuals, and ethical approvals were also secured for this aspect of the research. The tissue was collected in a specific medium and cut into small pieces. The pieces underwent digestion in a tailored solution and were processed further to create organoids, which were then placed into culture wells with a specialized growth medium.
Plasma proteomics
For protein isolation from plasma samples, cellular debris was removed through centrifugation. The clear supernatant was transferred for protein quantification using a standardized kit. High-abundance proteins were removed before the samples were treated for digestion, and then trypsin was added for an overnight reaction. The resulting peptides were prepared for mass spectrometry analysis.
During liquid chromatography and mass spectrometry, peptides were separated and analyzed for their respective mass, allowing for comprehensive identification through specific methods and careful adjustments at various stages of the process.
Missing value imputation
To manage any missing data points in our analysis, we applied a method known as K-nearest neighbours, which was implemented using Python’s scikit-learn functions.
Genetically engineered mouse models
We utilized several genetically modified mouse models, including ones designed with specific mutations relevant to pancreatic cancer. These models were generated and bred to achieve desired genetic backgrounds, with genotyping confirming the alterations. We also used tamoxifen for inducing specific genetic changes at set times.
Mouse blood samples and ELISA
Plasma from mouse blood samples was collected post-euthanasia, centrifuged, and processed to determine protein concentration. An ELISA was performed to measure specific protein levels based on established protocols.
Subcutaneous transplantation with mouse pancreatic cancer organoids
Immunocompromised mice were used for the transplantation experiments, where cells from organoids were injected under the skin using a gel-like medium. Various cell lines were tested, and different treatments like tamoxifen were applied based on the models set up. Tumor growth was closely monitored, with humane endpoints established to ensure ethical compliance.
Ultrasound-guided pancreas orthotopic transplantation
We adapted existing techniques for injecting pancreatic cancer cells directly into the pancreas of mice using ultrasound guidance. Careful positioning allowed for accurate injection, which was confirmed through imaging techniques.
Ultrasound imaging of pancreatic tumors
Ultrasound imaging was employed to capture detailed images of pancreatic tumors in the mice, which were carefully monitored to ensure the animals remained stable throughout the process.
Patient-derived organoid pancreatic cancer xenografts
Organoids were prepared for injection and tracked for tumor growth in a mouse model, with sizes recorded and tissues collected for further analysis post-experiment.
SPP1 neutralization experiments
In mice, SPP1 antibody treatments were administered to assess their impact on pancreatic cancer over several weeks.
Mouse tumor cell isolation and organoid cultures
Post-euthanasia, pancreatic tumors were processed to extract cells, which were cultured in controlled conditions. The initial cultivation emphasized selecting for cancer cells, and after some passages, the full growth medium was utilized.
Organoid staining
Organoids underwent a thorough staining process to visualize specific cellular markers, utilizing fluorescent techniques for assessment under microscopy.
Immunohistochemical staining
Following standard dissection, tumor samples were fixed and prepared for histological analysis, employing conventional staining methods to elucidate qualitative characteristics.
Immunofluorescence staining
Tumor tissue sections were prepared for immunofluorescence, with blocking and staining protocols established to visualize different antigens effectively.
CRISPR–Cas9-mediated knockout of target gene in organoids
Using the CRISPR-Cas9 system, specific genes were targeted and knocked out in organoids, following an established electroporation method for efficient genetic modification.
Flow cytometry
Cells from organoids were prepared and analyzed through flow cytometry to assess various characteristics, applying quality control measures during the process.
RT–qPCR
RNA was isolated from organoids for analysis, following specific treatments to gauge responses. Quantitative PCR was utilized to evaluate expression levels against housekeeping genes.
Western blotting
Cell lysates from treated cells were analyzed through Western blotting using established protocols for protein detection and quantification, ensuring compliance with standard practices.
Lentiviral production and infection
Lentiviruses were produced to facilitate the introduction of genetic material into organoids, with careful transfection and selection processes followed to ensure successful incorporation.
Chromatin immunoprecipitation
Chromatin immunoprecipitation was performed to evaluate potential transcription factor interactions with targeted gene promoters, utilizing established protocols for accuracy.
RNA-seq analysis
RNA sequencing was carried out to assess differences between various organoid samples, with extensive data processing for alignment and normalization applied throughout.
Gene set enrichment analysis
Analyses were conducted to understand gene expression patterns, employing specific methodologies to assess significance in relevant datasets from pancreatic cancer samples.
scRNA-seq analysis
Single-cell RNA sequencing data were downloaded, processed for quality, and analyzed for insights into cellular diversity within pancreatic ductal adenocarcinoma specimens.
Graphical abstracts
Diagrams for the study were created using design software, tapping into graphical tools to illustrate key findings succinctly.
Statistical analysis
All statistical methods were clearly defined, with appropriate software utilized to analyze data from different experiments consistently.
Survival analysis
Survival analyses were performed, based on comprehensive datasets from pancreatic cancer cases, to explore the impact of various treatments on overall outcomes.
Reporting summary
Additional details on the study’s design can be accessed in the linked reporting summary for further insights into the methodology and findings.