Participants
This study included 11 independent datasets with a total of 863 participants after quality control measures were implemented. Among these datasets were: (1) the PIPD dataset: 166 patients with Parkinson’s Disease (PD) and 60 healthy controls (HC); (2) the DBS-fMRI dataset: 14 PD patients assessed before and after DBS surgery and 25 HC; (3) the TMS dataset: 36 PD patients; (4) the DBS-SS dataset: 342 PD patients; (5) the DBS-ECoG dataset: 17 PD patients who underwent STN-DBS surgery; (6) the MRgFUS dataset: 10 patients with tremor-dominant PD; (7) the aDBS dataset: 4 PD patients; (8) the LCT dataset: 21 PD patients; (9) the ET dataset: 45 patients with Essential Tremor (ET) and 45 HC from the PIPD dataset; (10) the dystonia dataset: 42 dystonia patients and 21 HC; and (11) the ALS dataset: 30 patients with ALS and 30 HC. PD diagnosis followed the revised criteria from the International Movement Disorder Society (MDS, 2015) or the Chinese Parkinson’s Disease Diagnostic Criteria (2016 version). The sections below give detailed descriptions of each dataset.
PIPD dataset
Patients
From the Henan Provincial People’s Hospital in China, 180 patients with PD were enrolled based on certain criteria. Participants needed to be at least 18 years old and diagnosed with PD. Exclusion criteria included: (1) MRI contraindications; (2) prior neurological issues aside from PD; (3) any history of invasive neurosurgeries like DBS; and (4) considerable head motion during rsfMRI scans. After processing, 166 patients remained in the analysis (64 women, 102 men; average age 61.8 years). More demographic and clinical details are available in the extended data.
HC participants
A total of 71 healthy participants over 18 years old, with no neurological or psychiatric disorders, were recruited. Exclusion criteria were similar to the patient group. Following the removal of 11 participants due to excessive head motion, 60 HC individuals were included (34 women, 26 men; average age 56.10 years). Notably, demographic differences between the PD and control groups prompted matching of a subset of 65 PD patients for the analysis. The experiment was approved by the local Institutional Review Board (IRB), and all participants provided written informed consent.
MRI acquisition
The participants underwent one structural MRI scan lasting 8 minutes and 50 seconds, along with five scans of approximately 6 minutes and 14 seconds each, totaling 31 minutes and 10 seconds. The Siemens 3T Prisma MRI scanner was utilized, with structural scans performed using a T1-weighted MP2RAGE sequence. Specific details of the imaging parameters are provided in the original data.
DBS-fMRI dataset
Patients
This dataset included 14 PD patients diagnosed with akinetic-rigid dominant PD, recruited from various centers in China. The ethical approval was secured from multiple organizations, and each patient provided written consent. Inclusion criteria required participants to be aged 18-75, have specific clinical ratings, and a documented positive response to dopaminergic medications. Exclusions were broader, including individuals unable to follow instructions or those with relevant medical complications. Of the initial group, 11 patients provided complete data.
HC participants
HC participants of comparable age were enrolled, with similar exclusion criteria applied. Ultimately, the control group consisted of 25 individuals, with further details on their demographics and imaging included in supplementary data.
MRI acquisition
Data acquisition spanned multiple visits with distinct imaging sessions, both pre and post-surgery. The collection involved a variety of scans at different follow-up intervals, originally planned to gather comprehensive imaging and functional data on the participants.
TMS dataset
Patients
This dataset involved patients recruited between May 2023 and April 2024, with ethical oversight from the local IRB. Participation required a confirmed PD diagnosis and specific criteria that included being stable on medication. A total of 36 patients were randomly assigned to different treatment groups and received targeted TMS across fourteen consecutive days. Detailed methodologies for the assignment and group differentiation were rigorously followed, and the efficacy of stimulation was assessed over time.
MRI acquisition
Using the same MRI protocols as previously mentioned, scans were collected from each participant pre- and post-treatment.
DBS-ECoG dataset
Patients
Seventeen PD candidates for STN-DBS surgery were recruited, with all ethical protocols adhered to and informed consent obtained. Preoperative medication was paused, and ECoG arrays were used to gather high-resolution data on cortical activity during stimulation.
MRgFUS dataset
Patients
A group of 10 patients with tremor-dominant PD was enrolled to evaluate MRgFUS treatment, following the ethical guidelines and consent protocols. The methodology revolved around specific anatomical targets based on imaging criteria for effective lesioning.
DBS-SS dataset
Extracting DBS sweet spots involved a wide cohort and established methodologies to derive effective targeting regions based on past patient outcomes documented across a broad timeline. This aimed to refine stimulation techniques to optimize clinical results.
Clinical assessments
Motor symptoms were evaluated using various established scales, ensuring inter-rater reliability and consistency across assessments. Protocols across datasets followed rigorous ethical considerations and standardized clinical evaluations.
MRI preprocessing
The MRI data underwent a systematic preprocessing sequence aimed at enhancing analysis accuracy, accounting for motion artifacts and other physiological markers while standardizing the images for comparison.
RSFC analyses
Different types of analyses were conducted to explore connectivity patterns among various brain regions, utilizing seed-based methods to highlight significant neural pathways relevant to the conditions being studied.
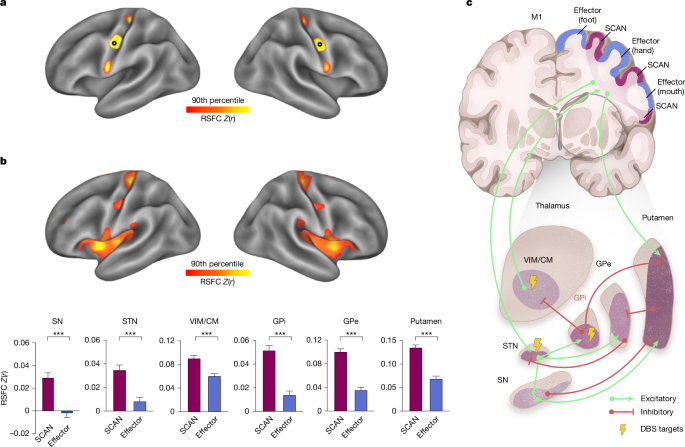
Identification of cortical SCAN regions
A two-stage analysis method was employed to delineate key cortical SCAN regions, leveraging both individualized and group-level data to understand functional connectivity throughout the Cortex.
Reporting summary
Detailed research design and methodology are accessible through supplementary materials linked to this article, aiming to foster transparency and reproducibility in scientific research.