Overview of Study Recruitment
This study gathered participants from the 23andMe customer network. They willingly provided informed consent and signed up for the research online, following a protocol sanctioned by the Salus Institutional Review Board, which is accredited by the Association for the Accreditation of Human Research Protection Programs. Only participants with verified consent were included in the analysis once data analyses commenced.
The 23andMe GLP1 survey started in August 2024 and sought to understand individuals’ experiences with medications known as GLP1 receptor agonists. It specifically targeted those who had indicated in a previous survey that they had used prescription medications for weight loss. Questions covered aspects like drug brand, dosing schedules, duration of treatment, effectiveness (including pre-treatment and current weight), side effects, and reasons for starting or discontinuing GLP1 treatment. The focus was primarily on six specific drugs: Ozempic, Wegovy, compounded semaglutide, Mounjaro, Zepbound, and compounded tirzepatide. The first three are variations of semaglutide, while the latter three relate to tirzepatide. A comprehensive list of survey questions can be found in Supplementary Table 24.
Phenotype Definitions
Using responses from the surveys, phenotypes were defined to capture various aspects of the drugs’ effectiveness and side effects. The efficacy phenotype was characterized by the change in BMI from before to after treatment (or the current BMI for ongoing treatments). For participants who had taken more than one GLP1 medication, the drug used for the longest duration was selected. Specifically, we calculated a percentage BMI change phenotype as follows:
$$\Delta BMI\% = 100(\mathrm{BMI}_2 – \mathrm{BMI}_1) / \mathrm{BMI}_1$$
In this formula, BMI1 and BMI2 indicate the participants’ pre-treatment and post-treatment BMI, respectively, where weight is measured in kilograms per height in meters squared. We applied quality checks, excluding anyone with a weight under 36 kg or over 181 kg, height below 1.39 m or above 2.06 m, BMI less than 14 kg/m2 or greater than 70 kg/m2, or age under 18. These filters eliminated 80 participants (0.29%). Further inspection of the ΔBMI% phenotype showed a heavily skewed distribution, leading to additional quality control that removed extreme outlier participants who experienced BMI changes beyond 20% or below -45%. The ΔBMI% values were marked as missing for those who didn’t pass quality control.
To facilitate genetic associations in weight units rather than ΔBMI%, we also identified a corresponding Δweight phenotype, reflecting the change in weight from baseline in kilograms. It’s important to note that, since adult height is treated as constant during the treatment, the percentage changes in BMI and weight are mathematically equivalent.
As for side effects, separate case-control phenotypes were defined for each side effect reported in the survey, comparing participants who rated their side effects as moderate or severe (cases) with those who felt their side effects were mild or absent (controls). Again, for participants using multiple GLP1 medications, the one used longest was prioritized.
We also created phenotypes representing covariates for drug type (semaglutide = 1 versus tirzepatide = 0), dosage, and duration of treatment. The dosage was based on the most recent reported weekly amount in milligrams, whether it was the final or current dose for ongoing users.
Comparison of Self-Report and EHR Data
As a part of the 23andMe experience, participants can share electronic health record (EHR) information collected through their Apple iPhone devices. Specifically, the Apple Health application allows integration with healthcare providers for sharing EHR data with third parties via Apple HealthKit. Participants can choose to share these records for research purposes, and we utilized this information to compare it with the self-reported survey data. Detailed comparison methods can be found in Supplementary Information.
Non-Genetic Predictors of BMI Loss
To investigate how non-genetic factors affect BMI loss, such as drug type, dosage, and treatment duration, we constructed the following model:
$$\Delta BMI\% \sim \mathrm{age} + \mathrm{sex} + \mathrm{BMI}_1 + \mathrm{drugType} + \mathrm{dose} + \mathrm{days}_{\mathrm{treat}} + \mathrm{drugType} : \mathrm{dose} + \mathrm{drugType} : \mathrm{days}_{\mathrm{treat}} + \mathrm{dose} : \mathrm{days}_{\mathrm{treat}} + \mathrm{dose} : \mathrm{days}_{\mathrm{treat}} : \mathrm{drugType}$$
In this equation, ‘drugType’ signifies whether semaglutide (1) or tirzepatide (0) is used, ‘dose’ indicates the dosage in milligrams, and daystreat represents the total treatment days for the respective drug. The colon (:) denotes interaction terms among variables. Notably, semaglutide and tirzepatide come with varying standard dosages, which the regression model accounts for with the drugType:dose interaction term.
Genotyping and SNP Imputation
DNA extraction and genotyping took place using saliva samples in labs certified under Clinical Laboratory Improvement Amendments and accredited by the College of American Pathologists. The genotyping was done on various platforms. For instance, the V1 and V2 platforms (Illumina HumanHap550 BeadChip) typed approximately 560,000 SNPs, including around 25,000 custom SNPs from 23andMe. The V3 platform utilized the Illumina OmniExpress BeadChip with close to 950,000 SNPs, with custom content added for better compatibility with the V2 array. The V4 platform was an entirely customized array of approximately the same number of SNPs, featuring reduced redundancy from V2 and V3 SNPs while including more coverage of rarer coding variations. Lastly, the V5 platform was based on the Illumina Global Screening Array, with roughly 654,000 preselected SNPs and about 50,000 custom variants. Genotype data for participants were compared against a reference panel from the Haplotype Reference Consortium, enhanced with additional sequences for improved imputation performance and detailed further in Supplementary Information.
Association Testing
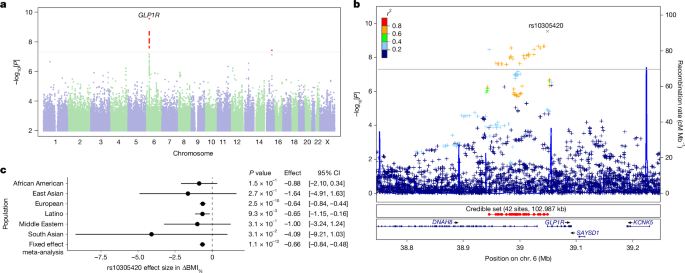
A GWAS evaluating ΔBMI% was conducted among individuals of European ancestry utilizing previously described methods. Essentially, unrelated participants were included based on their genetic ancestry groups. The GWAS, along with covariates outlined earlier, also integrated five genetic principal components to refine genetic ancestry and included variables to accommodate variations in genotyping platforms. From a total of 21,822 participants of European ancestry, we required complete data to construct the necessary target phenotype, resulting in 18,488 qualified participants. We filtered participants by relatedness, ensuring no two individuals shared more than 700 cM identity by descent, akin to sharing between first cousins and yielding a final GWAS sample of 15,237. A similar approach was followed for GWAS on side effect phenotypes. For investigating drug-specific associations, we ran the GWAS separately for those treated with semaglutide and tirzepatide, eliminating the drug-type variable and related interaction terms when needed. All GWAS results were corrected for inflation, with no inflation factor exceeding 1.035 across phenotypes.
We opted not to perform testing in non-European populations due to smaller sample sizes. Instead, the focus was on variants found associated in the European GWAS, checking for associations in non-European cohorts using similar methods.
Replication
Replication of the efficacy associations identified took place in the All of Us cohort, extracting genomic and EHR data along with a drug code for either semaglutide or tirzepatide from 9,579 participants. After filtering for relevant information regarding pre-treatment and post-treatment BMI and genotype data passing quality checks, a total of 4,889 participants were included, 3,948 of whom had complete data when incorporating relevant covariates. The replication analysis examined associations between the EHR-derived ΔBMI% and the genotype, including covariates. We also conducted the analysis with mean-imputed missing drug dose data, allowing for a larger sample size of 4,855.
Furthermore, we attempted a replication analysis within the UK Biobank cohort, but the data available did not align with the use of semaglutide or tirzepatide, focusing instead on earlier GLP1 receptor agonist variants. Methodology details for this replication analysis are available in Supplementary Information.
Genetic and Non-Genetic Risk Modelling
To create a collaborative model for ΔBMI% and treatment-related side effects, we used a selection of treatment and clinical variables alongside demographic and genetic factors. In addition to variables in the GWAS, years of education as a socio-economic status proxy and binary indicators for previous diagnoses (T2D, hypertension, non-alcoholic fatty liver disease) were included. Continuous predictor variables were standardized prior to modeling for effect size comparison.
A linear multi-variable model was applied to fit ΔBMI%, while logistic regression models suited the binary side effect definitions. The data were divided randomly into training (70%) and testing (30%) sets, with the test set assessing model performance. Further details can be found in Supplementary Information.
To evaluate the efficacy model’s performance, we applied it to a sample of 642 individuals who had provided HealthKit EHR data but did not take part in the GLP1 survey. In this scenario, we assumed unknown dose, treatment duration, and drug type variables, assigning arbitrary constant values across all participants.
Reporting Summary
More information on the research design can be found in the linked Nature Portfolio Reporting Summary related to this article.