Tissue acquisition for single-nucleus multiome profiling and spatial transcriptomics involved obtaining snap-frozen samples from decidual and basal plate tissues at Stanford University and the University of California, San Francisco. All samples were collected with written consent from women undergoing elective terminations of normal pregnancies or after term deliveries. The clinical records were reviewed to filter out any placenta-related complications.
Fresh placental tissues were examined and dissected under a microscope by pathologists. The decidua basalis was carefully micro-dissected on ice and identified based on its distinctive histological features. The dissected tissues were washed to remove any remaining blood cells and then flash-frozen in liquid nitrogen for later processing. Only samples with a RNA Integrity Number (RIN) of 7.0 or above were used for further analysis.
To isolate single nuclei from these tissues, the method involved grinding the samples on dry ice before homogenizing them in a specific buffer. After filtration, the nuclei were purified through a density gradient. Around 15,000 nuclei per sample were then processed using a specific platform for sequencing.
For CODEX imaging, placenta samples were embedded in OCT and sliced into thin sections for further processing. These sections underwent a series of treatments before being incubated with a cocktail of barcoded antibodies. The imaging system used a detailed protocol across multiple channels to capture the data, and the antibody panel was assembled using validated sources.
Single-nucleus Multiome libraries were constructed using a specific kit, where isolated nuclei were encapsulated and subsequently processed for gene expression and chromatin accessibility. High-quality nuclei profiles were retained based on set criteria, and analysis on open-chromatin peaks was conducted, integrating data across samples.
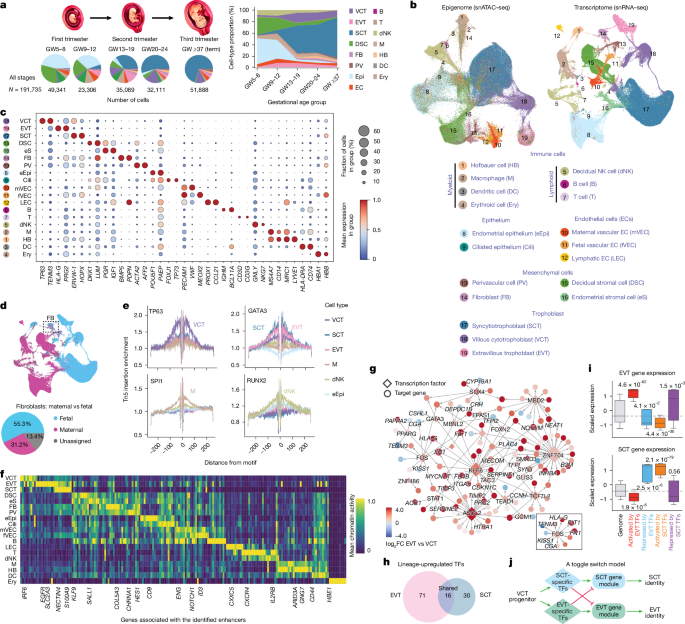
Cell type-specific gene regulatory networks were constructed using a workflow that combined chromatin accessibility with gene expression data. The analysis aimed to define transcription factor-target gene interactions across different lineages, resulting in a detailed overview of regulatory rewiring based on cell types.
Differential expression analysis compared various cell types, deriving significant data for further gene ontology analysis. Pseudotime trajectories were inferred for different cell populations to understand their developmental pathways.
Spatial transcriptomic analyses utilized a high-resolution sequencing platform designed to provide an atlas of gene expression directly from tissue sections. The preparation involved meticulous steps, including tissue permeabilization and in situ reverse transcription.
Communication among cells in spatial contexts was analyzed to explore intercellular signaling pathways. A supervised machine-learning model was employed to identify potential invasive characteristics of extravillous trophoblasts (EVTs) based on specific gene profiles.
Immunofluorescence techniques were used to visualize cellular components, while single-molecule fluorescence in situ hybridization was performed for precise localization of specific probes.
In vitro experiments observed decidualization of cells and their interaction with certain substances, alongside a comprehensive analysis of the resulting cellular pathways.
A Transwell invasion assay was set up to examine cell invasiveness in varying conditions, emphasizing the role of different treatments on trophoblast behavior.
The study also included a single-cell risk assessment related to pregnancy complications, integrating various datasets to identify cell types at risk. All human tissue samples used were collected following ethical guidelines, with necessary approvals obtained from institutional review boards.