Human Tissue Sample Collection and Compliance
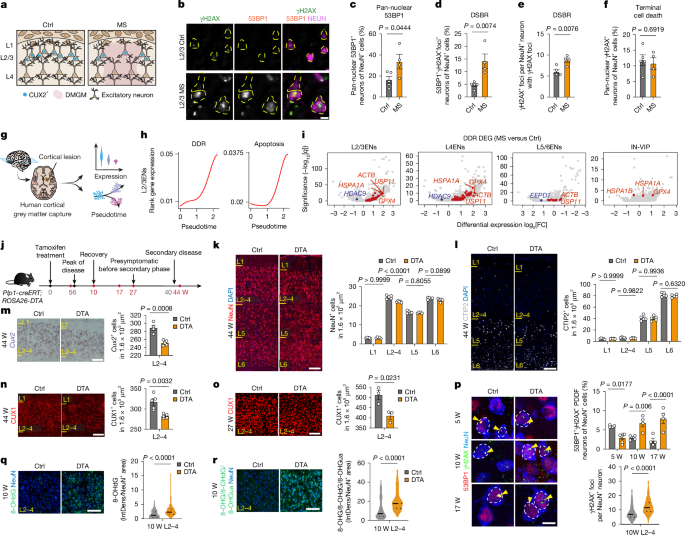
A previous study conducted single-nucleus capture and RNA sequencing analysis on age- and sex-matched human control and multiple sclerosis (MS) cortices. For this new research, post-mortem human brain samples showing MS grey matter pathology were sourced from the UK Multiple Sclerosis Tissue Bank based at Imperial College London. The National Research Ethics Committee in the UK granted ethical approval for utilizing these samples. In total, nine snap-frozen brain blocks were analyzed—comprising four from MS patients and five from individuals without neurological diagnoses—using immunohistochemistry techniques. For expression analyses of genes CUX2 and ATF4, a 25-year-old sample was sourced from the Pediatric Neuropathology Research Laboratory at the University of California, San Francisco, adhering to guidelines established by their Committee on Human Research and Institutional Review Board. Participant details can be found in the supplementary material.
Primary Rat Glial Cultures
Fresh postnatal day 5 Wistar/Han rat heads were obtained and kept in ice-cold Hibernate-A medium. Whole brains were extracted, homogenized, and centrifuged. The supernatant was discarded, and the remaining tissue was resuspended in a prepared dissociation solution that had been heated and shaken. After further centrifugation, the tissue was resuspended in a cold neutralizing solution. A series of triturations and rest periods were executed to create a single-cell suspension, which was filtered and subjected to isotonic Percoll density centrifugation. The cell pellets were processed and prepared for sorting with specific microbeads targeting CD11b/c markers, then microglia were collected and treated for growth in specific media formulations. Astrocytes were obtained by further sorting, subjected to incubation and specific media treatments until passaged and concentrated in conditions allowing for study.
Demyelination and Neuroinflammation Models
The study utilized ROSA26–eGFP-DTA mice bred with PLP/creER mice to create PLP/creER–ROSA26–eGFP-DTA mice. Guidelines set by the Animal Care and Use Committee were followed for all procedures. Mice were treated with tamoxifen, and brain tissues were collected at various intervals to compare the impact of the treatments. Additionally, Myrf and Sox10 strains were also crossed to generate mutant mice for further study under ethical oversight.
Ectopic IFNγ Expression Model
To investigate the role of IFNγ in central nervous system remyelination, double-transgenic mice were produced. These mice were maintained on a doxycycline-restricted diet until the age of eight weeks, at which point the suppression was lifted, and subsequent observations were made regarding the brain’s response, adhered to ethical standards.
KO and cKO Lines
Various mouse lines, including Cux2 and Atf4, were maintained under specific housing and care protocols following institutional guidelines. All interactions and experiments were documented for compliance and reproducibility.
Neonatal Hypoxia and Acute Neuroinflammation Model
Neonatal mice and their parents experienced a chronic hypoxia treatment aimed at simulating conditions that could affect healthy neuronal development. Following this, they were returned to normal oxygen levels prior to brain tissue collection, with all procedures thoroughly monitored and ethical approvals obtained.
Acute Demyelination with Cuprizone
Mice received intraperitoneal tamoxifen before a cuprizone diet was implemented to induce demyelination over several weeks, with daily monitoring and final brain tissue evaluation adhering to compliance standards.
Collection of Tissue for Histology
Immunohistochemistry methodologies were applied to perfused mice to collect brain tissues for analysis, which were later sectioned and preserved under specific conditions for study.
Collection of Tissue for snRNA-seq
For snRNA sequencing, brain cortices were collected in cold buffer solutions and stored at low temperatures until sequencing was performed.
snRNA-seq
RNA extraction from cell nuclei involved barcoding processes followed by sequencing using advanced platforms, and contamination measures were taken to ensure data integrity.
Cell Type Annotation
A combination of clustering and reference-based analytical techniques were utilized to annotate various cell types derived from the collected data, with validation against established databases.
DEG Analyses
Differentially expressed genes were identified through various statistical tests and analytical platforms, and visual representations were generated to illustrate findings.
Trajectory Pseudotime and Real-Time Analysis
Gene expression was analyzed over time across different conditions, applying mathematical models to assess variations in gene activity at specified intervals.
GO Analysis
Gene ontology analyses were conducted to uncover biological processes impacted by different treatments, utilizing computational tools for statistical evaluations.
DDR Gene List
A comprehensive list of genes related to DNA damage repair was compiled and refined following established protocols.
Quantitative PCR
RNA was extracted from culture cells with subsequent reverse transcription performed to analyze gene expressions, following specific procedures outlined by manufacturer instructions.
Neurogenin-2 Knock-in iPS Cells
Ribonucleoprotein complexes were developed for editing iPS cells, followed by selection and assessment methods to identify successful modifications.
Cell Lines and Culture
Various established cell lines were maintained under standard conditions, with each being validated for routine checks against contamination.
Induced Neuron Generation and Culture
Protocols for differentiating iPS cells into neurons were executed with precise attention to media conditions and timing aimed at promoting neuronal development.
Plasmid Construction, Lentivirus Preparation and Infection
Donor plasmids were constructed for viral expression systems, and procedures were put in place for efficient delivery into target cells.
Comet Assays
Comet assays were performed post-treatment to assess DNA damage, employing precise techniques for assessment and image acquisition.
MTT Viability Assay
Cytotoxicity and viability tests were conducted through established methods, with data recorded for subsequent statistical analysis.
CellROX Probe Assay
Cellular oxidative stress was analyzed utilizing specific probes and imaging techniques to observe and quantify responses.
Luciferase Assay
Promoter activity was assessed through dual-luciferase assays, ensuring controls were applied for accurate comparison.
NHEJ Reporter Assay
Cellular responses to DNA repair signals were examined using specific reporter assays, with results analyzed for efficiency of repair mechanisms.
Immunohistochemistry Analysis of Rodent Brain Tissue
Rodent brain tissues underwent a series of washes and staining to visualize targets of interest, utilizing specific antibodies for detection.
In Situ Hybridization on Rodent Brain Tissue
This method was employed to localize specific mRNA within the tissue, providing insights into gene expression patterns.
Immunohistochemistry Staining of Human Post-Mortem Brain Tissue
Methods were employed to detect specific protein expressions in human tissues, ensuring rigorous protocols were followed for staining and imaging.
Immunocytochemistry
Cell cultures were fixed and processed for antibody staining, with analyses focused on identifying specific cellular markers through visual techniques.
smFISH
Single-molecule fluorescence in situ hybridization techniques were adopted to investigate specific transcript localizations within cells.
Immunoblotting
Cells underwent lysis for protein extraction before quantification using standard protocols to determine protein presence and levels.
Imaging
Standard and confocal imaging techniques were applied to acquire data, ensuring all protocols adhered to for accurate representation.
Layer Thickness and Cortical Cell Counts
Thick sections of cerebral cortex were analyzed to measure layer thickness and cellular composition, maintaining strict methodological standards.
Quantification of DNA Damage Marks
Analysis of DNA damage markers was meticulously conducted across selected brain regions, with manual counts verified through stringent methods.
Study Design and Statistical Analysis
The rationale for sample sizes was derived from previous studies, aiming for a balance between statistical power and feasibility in experimental designs, with considerations given to demographic factors.
Materials Availability
Materials generated through this study are accessible via a formal request, contingent on completion of appropriate agreements.
Reporting Summary
Further design details regarding the research are included in the supplementary materials linked to the article.