Dataset Collection and Processing
Information on Case Detections in North America
In this study, a detection is defined as a positive PCR test from a collected sample. In Canada, year-round surveillance in both wild and domestic populations is coordinated by several agencies, including the Canadian Food Inspection Agency and the Public Health Agency of Canada. In the USA, the Animal and Plant Health Inspection Service (APHIS) oversees HPAI surveillance and testing in wild birds via reported morbidity and mortality events, hunter-collected waterfowl, and environmental sampling. They also monitor domestic birds using various reporting methods, including mandatory tests through the National Poultry Improvement Plan.
Data on HPAI detections in the USA for this analysis came from reports downloaded in November 2023. The majority of detections during the study period (November 2021 to September 2023) were in wild birds. Domestic bird detections were categorized by type and whether they were from commercial farms or backyard flocks, the latter being those with fewer than 1,000 birds. Most detections among domesticated species were reported from commercial chickens, turkeys, and other categories, including backyard birds.
This outbreak has also affected a variety of mammals, with detections reported in species like red foxes and skunks. Other hosts include a wide array, including seals and bobcats.
Genomic Data Processing and Initial Phylogenetics
We collected nucleotide sequencing data for HPAI clade 2.3.4.4b H5Nx viruses from the GISAID database. Sequences were aligned and visually inspected, followed by the removal of significant gaps. Deduplication of identical sequences was performed, and temporal outliers were cut out through initial phylogenetic reconstruction. This resulted in a dataset of 1,824 sequences for further analytical use.
Biases in Genomic Data and Ne Inference
North American sequencing data is primarily skewed towards the USA. To assess whether the sequencing data reflect case detections accurately, we inferred viral Ne, which is related to genetic diversity and disease prevalence. The inferred Ne suggested a positive correlation with detections, with increased Ne values appearing about a week before detection spikes.
Despite the uneven acquisition of sequences over time, they seem to roughly represent the H5N1 case amplitude. Given these insights, we used the full sampling period for broader geographical and introduction analyses, but focused on the first six months for more detailed transmission pattern reconstructions.
AVONET Database
We downloaded and merged data from the AVONET database with host metadata for each sequence. Taxonomic details for host species were used for further merging, aiming to ensure accuracy in ecological data. Domesticity status was determined based on available metadata.
Phylodynamic Analysis
Bayesian phylogenetic reconstructions were conducted, using specific models for each dataset. Four independent MCMC chains controlled for valid sample sizes and reasonable estimates were run. The outputs were combined and subjected to further analyses.
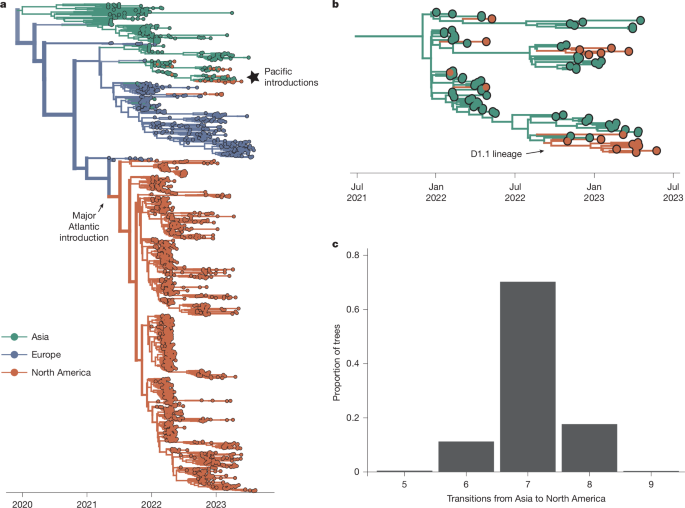
Geographical Introductions Analysis
This analysis characterized HPAI introductions in North America by sampling sequences from Europe and Asia while also accounting for all available North American sequences. The USA sourced the majority of samples, with Canada and a few Central American countries contributing relatively small numbers.
Migratory Behavior Analysis
We defined discrete traits for our modelling based on metadata, categorizing birds by migratory behavior and creating a subsample focused on migratory traits. The selection aimed for balanced representation across behaviors.
Domestic/Wild Titration Analysis
To assess how the sampling of wild birds influenced the estimation of rates between domestic and wild birds, we performed multiple datasets with varying wild sequencing ratios, culminating in a final dataset that provided insight into the dynamics of domestic versus wild populations.
Assessment of Sampling Bias
To determine the correlation between discrete traits and ancestry, we employed various association tests and sensitivity analyses to understand the implications of sampling biases on our findings.
Our analysis endeavors to refine insights into HPAI dynamics based on robust genetic data while acknowledging potential discrepancies arising from sampling methods.