Clinical Samples and Cohorts
Supplementary Table 10 provides an overview of the clinical cohorts discussed in this article, along with their analysis results. Detailed descriptions of each cohort can be found in the text and again in Supplementary Table 10.
Neoadjuvant Anti-PD-1 cSCC Clinical Trial Cohorts
The protocols for the clinical trials, alongside the statistical analysis plans and institutional review board approvals, have been documented previously. To summarize briefly, the trials adhered to the principles set forth in the Declaration of Helsinki and the International Conference on Harmonization Good Clinical Practice guidelines. All participants provided written informed consent, and the authors had unrestricted access to the data, bearing full responsibility for its content. Patients aged 18 and older with resectable stage II–IV (M0) CSCC were included, with primary surgery recommended as per standard clinical practice. For stage II patients, a primary tumor diameter of at least 3 cm was required. Patients also needed adequate organ function, assessed through complete blood count and metabolic function tests, at least one measurable lesion according to the RECIST criteria, and an ECOG performance status of 0 or 1. Following a screening period of up to 28 days, patients received neoadjuvant cemiplimab (Regeneron Pharmaceuticals) at a dosage of 350 mg intravenously every 3 weeks, continuing until unacceptable toxicity, disease progression, or withdrawal of consent occurred. Imaging assessments were conducted at baseline, and at weeks 6 and 12. Upon completion of the neoadjuvant treatment, surgery was scheduled between study days 75 and 100. If early discontinuation of cemiplimab occurred, the treating physician had the option to advance surgery. Biopsy samples collected prior to treatment underwent histopathological assessment to confirm diagnosis and allow for morphological comparisons of tumor tissue before and after therapy.
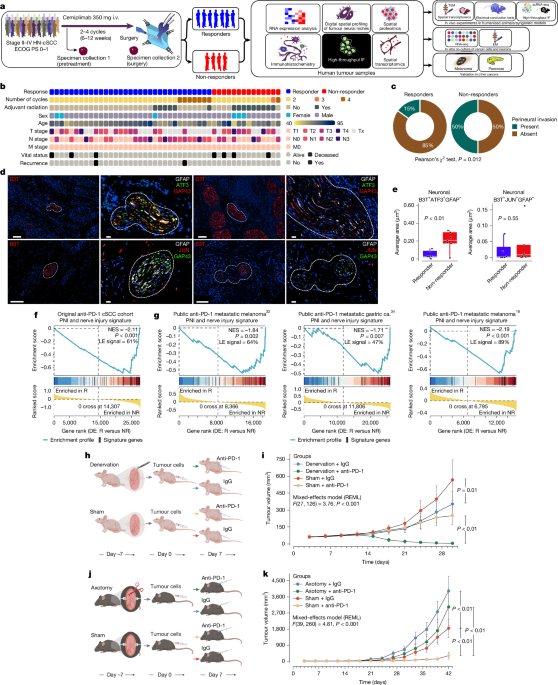
Pathologic response assessments in post-treatment surgical specimens utilized standard evaluation recommendations and were reviewed by a dedicated dermatopathologist to ensure standardized reporting. The primary endpoint was defined as the pathologic complete response, characterized by the absence of viable tumor in post-treatment specimens, determined through independent central pathology reviews. Major pathologic response, a secondary endpoint, required the presence of 0% to 10% viable tumor cells in these post-treatment samples. The specified definitions for these responses adhered to immune-related pathology criteria. Further details about the trial patients can be viewed in Fig. 1b, and the allocation of patient tumor samples to various analyses is outlined in Extended Data Fig. 1.
Immunohistochemistry Analysis of cSCC Anti-PD-1 Clinical Trial Samples
Immunohistochemical analysis was conducted on 4-μm-thick formalin-fixed and paraffin-embedded cSCC tissue samples collected pre- and post-anti-PD-1 therapy. Staining utilized antibodies against CD8, PD-1, and PD-L1, employing a BOND-RX instrument with a detection kit. Stained slides underwent scanning and digitalization via a Scan Scope XT system. A pathologist conducted single-stain annotations, with staining quantification performed using dedicated software to calculate marker-positive cell density. Statistical analysis applied two-tailed Mann–Whitney U-tests.
High-Plex, High-Dimensional IF Staining and Analysis
The Lunaphore Comet was employed to stain and image the FFPE samples. Slides were subjected to a microwave treatment to achieve dewaxing and rehydration, followed by antigen retrieval. Post-treatment, samples were cooled and placed in the Comet system for staining and imaging. Antibody panels were optimized through a characterization protocol to ensure optimal signal detection. Four specific mIF panels were utilized for patient tissues, with two panels applied to mouse tissues. The Comet collected images that were analyzed using a specific imaging software. A deep learning classifier assisted in identifying regions of interest within the tissues.
GeoMx DSP Experimental Design
FFPE tumor samples from the cSCC anti-PD-1 clinical trial were used for profiling neural and immune protein expression. High-resolution imaging was conducted, along with detailed mapping of neural niches. Viable tumor areas and perineural microenvironments were assessed using a combination of immuno-oncology protein assays. The GeoMx DSP software facilitated information visualization and statistical analysis, supporting the identification of protein expression patterns.
Bulk RNA-Sequencing of cSCC Anti-PD-1 Clinical Trial Tumor Samples
RNA extraction and library preparation followed standardized protocols, with quality assessments conducted on the isolates. Sequencing was carried out using a NovaSeq system. The resulting data was thoroughly examined for quality and aligned to a reference genome. Statistical evaluation provided insights into gene expression differences, which were then deposited in the NCBI database.
PNI and Nerve Injury Signature
The PNI signature utilized in this analysis was built from previously published gene expression signatures associated with PNI in cSCC. A comprehensive analysis culminated in the establishment of a gene set enriched in PNI-positive cases, initially tested using GSEA on the anti-PD-1 trial data to ascertain its enrichment in pretreatment tumor samples.
Original Clinical Cohorts and Non-Trial Samples
Beyond the trial-specific data, original clinical cohorts included treatment-naive patients with stage I–II cSCC and PDAC, whose tumors were subjected to extensive analysis. The findings contributed to wider examinations of immune infiltration and other relevant metrics.
Spatial Transcriptomic Analysis
Utilizing fresh-frozen and FFPE archived samples, this analysis sought to explore the spatial relationships between nerve injury and immune phenotypes, employing state-of-the-art sequencing technologies. Tissue sections underwent detailed processing to prepare libraries for efficient sequencing and analysis.
In Vitro and In Vivo Studies
The article describes a series of in vitro experiments with human and mouse neurons, alongside various assays to assess the impact of treatment on neuronal health. Co-culture systems with cancer cells allowed for exploration of interactions between neuronal activity and therapeutic responses. Detailed methodologies ensured the robustness of findings across various experimental conditions.
Statistical Analysis
Analyses employed a mix of statistical methods to assess data from in vitro and in vivo experiments. The significance of different variables was evaluated, ensuring reliable conclusions could be drawn. All data obtained are meticulously documented and shared for further scrutiny.