Mice
The study’s experiments were reviewed and approved by either the MIT Committee on Animal Care or the MGH Institutional Animal Care and Use Committee. For the research, female B6 and NSG mice aged between six to ten weeks were utilized. The animals were kept at a comfortable temperature (between 18–23°C) and humidity (40–60%), on a standard 12-hour light-dark cycle, with free access to food and water. All groups were age-matched and randomly assigned to either treatment or control groups to reduce bias. Randomization was implemented through a random numbering scheme right at the point of injection or treatment. It’s worth mentioning that researchers were not blinded during the injections or treatment phase, although they were blinded during data collection and quantification. There was no statistical method to determine sample size beforehand. Animal studies were conducted once for each condition unless stated otherwise in the figure legends. The data collected represented distinct animals, indicating biological independence. Mice were humanely euthanized when tumor size reached 2 cm or sooner if they exhibited criteria for humane endpoints, such as significant weight loss or signs of distress. Importantly, these institutional limits were maintained throughout all experiments.
Cell lines and culture
The human cell lines for this study were sourced from the American Type Culture Collection—specifically, HCC1806 (CRL-2335) from a female patient, and MDA-MB-231 (HTB-26) from another female patient. Mouse EO771 breast cancer cells came from CH3 BioSystems. The credibility of these cell lines was authenticated by the suppliers; the authors did not carry out any additional validation. Regular tests for mycoplasma contamination were conducted using the MycoAlert PLUS Detection Kit. All cells were grown in a Heracell humidified incubator, at 37°C with 5% CO2. The cell lines were typically maintained in RPMI-1640 medium that was supplemented with 10% heat-inactivated fetal bovine serum. However, for certain experiments, 10% dialyzed fetal bovine serum was used instead.
A modified version of RPMI-1640 medium was created, omitting serine or arginine, based on prior methodologies. Essentially, sufficient amounts of all amino acids and sodium phosphate dibasic were weighed out to prepare 25 liters of medium, followed by homogenization using a coffee grinder cleaned with methanol and water. Afterward, the resulting powders were mixed with water along with sodium bicarbonate and the base RPMI-1640 medium. Serine or arginine was prepared separately in water and added back to achieve standard concentrations.
Similarly, DMEM medium without serine or arginine was produced using a base powder formulation that lacked all amino acids and pyruvate. To this base, glucose, sodium pyruvate, and sodium bicarbonate were incorporated to match the standard DMEM composition. Amino acids were processed like in the RPMI method, ensuring the base was reconstituted properly.
Isolation of interstitial fluid, CSF and plasma
Interstitial fluid was extracted from mouse tissues using a technique adapted from established protocols. Female NSG or B6 mice aged 6 to 9 weeks, fed ad libitum, were used for collecting interstitial fluid samples. For each sample, organs from five mice were pooled together and treated as separate data points during analysis. Mice were euthanized consistently at the same time of day for metabolic consistency—this was done through cervical dislocation to avoid metabolic issues from CO2 exposure. Tissues remained chilled during the harvesting process. Once prepared, they were briefly rinsed with ice-cold saline, excess moisture was absorbed with filter paper, and the tissues were placed into conical vials fitted with nylon mesh filters. The samples underwent centrifugation, and the flow-through was collected and frozen for future use. Plasma was collected through cheek bleed and similarly processed before being pooled for analysis.
Cerebrospinal fluid (CSF) was obtained from the brains of the mice via established methods. In brief, the mice were anesthetized with a ketamine-xylazine combination, and CSF was extracted through the cisterna magna. Blood was also collected following CSF extraction, utilizing cardiac puncture, and the processing was completed expeditiously to preserve sample integrity.
In a separate experiment focused on the stability of metabolite levels, blood and liver interstitial fluid samples were either processed immediately or held on ice for varying durations before analysis. Subsequent measures quantified changes based on time.
MFP and brain tumour model generation
MDA-MB-231, HCC1806, and EO771 cell lines were modified with a lentiviral vector that included Gluc and GFP sequences. The top 10% of GFP-positive cells were then isolated via FACS. To create MFP (mammary fat pad) tumors, the mice underwent anesthesia before receiving injections of the cancer cells. For brain tumors, a similar procedure was followed, with careful injections performed into the specified point of the mouse brain. Tumor sizes were evaluated regularly through blood sampling and luminescence measurement using specific equipment.
Intracardiac injection and metastasis quantification
The same set of cancer cell lines was used, modified again with a lentiviral vector showcasing Fluc and GFP, and subsequently sorted to isolate the GFP-positive cells. Intracardiac injections involved injecting a specified number of cells into the heart under anesthesia, guided through ultrasound technology. Metastasis was monitored using bioluminescence imaging (BLI) systems, with protocols established for monitoring the luminescence at regular intervals.
Upon reaching the experiment endpoint, tissue samples were gathered for thorough analysis, with immediate imaging done post-euthanasia.
Generation of metastatic cell lines
After conducting intracardiac injections and BLI, any resulting GFP-positive tumors found in the brain or liver were isolated for further study. The tissues underwent a digestion process to facilitate breakdown, followed by cell extraction. These cells were then processed and expanded in culture, with GFP-positive populations isolated for future analysis.
Petal plot generation
To account for initial activity and variations across different tissues in the luciferase data, normalization procedures were employed against control lines. This was done through a statistical bootstrap approach that allowed for estimating mean values and their confidence intervals. Values indicating increases were capped at a given point, while the resulting data visualization was inspired by existing methodologies.
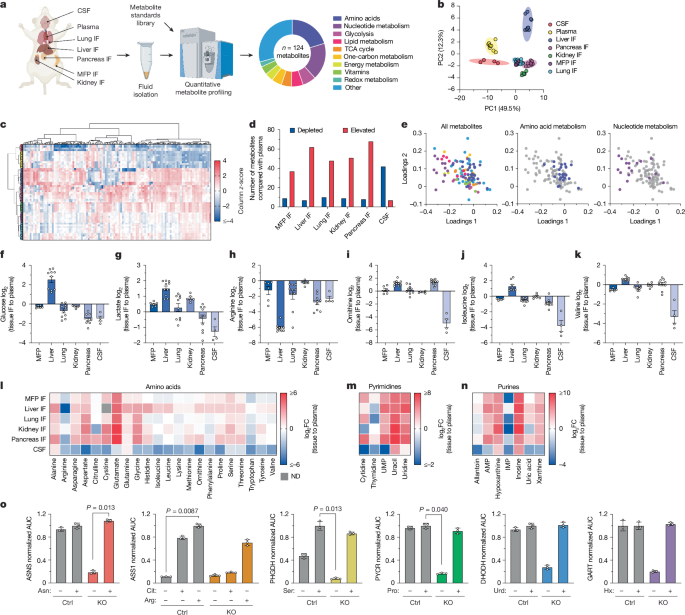
Quantification of metabolite levels in biological fluids
The quantification process for metabolites in mouse fluid samples followed established techniques. Typically, small sample volumes were mixed with a specific solvent to facilitate extraction. Various operational steps were taken, ensuring that the metabolites identified could be analyzed efficiently using high-performance liquid chromatography coupled with mass spectrometry.
Dietary serine or glycine restriction and metastasis assay
To examine how nutritional availability impacts metastasis, female NSG mice were placed on either a control diet or a high-serine diet before undergoing cancer cell injections. Upon completion of the dietary phase, tissues and plasma were collected for metabolomic analysis, ensuring all measurements adhered to consistency regarding time of day and methodology.
In vitro [U-¹³C]-glucose tracing
For in vitro studies, MDA-MB-231 cells were set up with specific methodologies to trace glucose metabolism. These cells underwent meticulous washing and subsequent treatment with glucose labeled with a specific isotope before being prepared for extensive analysis.
LC–MS analysis
Metabolite profiling was achieved using advanced mass spectrometry techniques, coupled with high-performance liquid chromatography. Standard calibration procedures were regularly followed to maintain accuracy in readings during the various phases of sampling.
Cell proliferation
To monitor cell growth over time, live-cell imaging techniques were employed, paired with metabolic supplementation as needed depending on the experimental conditions set forth by the study’s design.
Western blot
This technique was utilized to analyze protein expression in cells post-extraction. Various steps were taken to ensure samples were prepared appropriately before being subjected to electrophoresis and subsequent detection protocols.
Lentivirus generation
Procedures for producing lentiviral particles incorporated established transfection and collection methodologies, allowing for future experimentation on target cell lines.
CRISPR–Cas9-mediated gene knockout
For specific gene targeting, a series of rigorous processes were followed, including strategic RNA design and selection. Subsequent categorization of cells was done to isolate successful gene knockout events for further research into cellular behavior in the context of cancer.
DepMap and metastatic potential data
Gene expression data were aggregated from established online resources and utilized for correlation analyses regarding metastatic potential across various cancer cell lines.
Statistics and reproducibility
All experiments were performed with careful attention to reproducibility and statistical integrity, with multiple independent validations from in vitro to animal testing phases documented accordingly throughout the study.
Illustrations
Graphical representations relevant to the experimental design and findings were visually articulated using BioRender.
Reporting summary
Details regarding research design can be found in the linked reporting summary associated with this article.