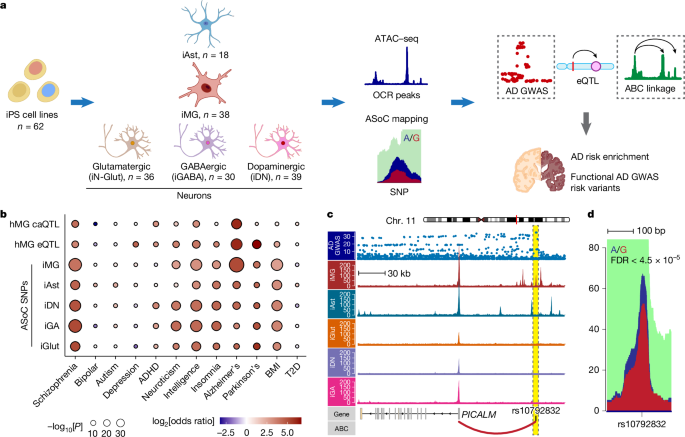
Human iPS cell lines and culture
The human iPS cell lines for ATAC-seq were created at the Rutgers University Cell and DNA Repository (RUCDR). These cells were generated using the Sendai virus method, which is crucial for preventing integration issues. A range of quality-control measures were put in place—this included immunofluorescence staining for pluripotency, tests for mycoplasma contamination, and in-house RNA-seq-based pluripotency testing (Pluritest). Additionally, eSNP-karyotyping or G-band karyotyping was performed at RUCDR. All donors were of European descent and their samples had been previously used in schizophrenia GWAS studies. No large copy-number variants (>100 kb) were found in any of the donors. The cohort included 29 schizophrenia cases and 33 control samples, comprising 37 males with an average age of 49.5 years, though the specifics about case-control status or age did not influence ASoC mapping results. Two human iPS cell lines obtained from control donors, homozygous for APOE3 (designated CD04 and CD09), were utilized for CRISPR–Cas9 editing. The human iPS cells were cultured using a feeder-free protocol on Matrigel-coated plates and maintained in mTeSR plus medium. Media changes occurred every second day, and cells were passaged as clumps every 4–6 days. All cultures were tested for mycoplasma contamination. Human iPS cell lines were sourced from the RUCDR NIMH Stem Cell Center, derived from cryopreserved lymphoblasts collected by the Molecular Genetics of Schizophrenia (MGS) consortium, under informed consent, approved by the Endeavor Health IRB, which also sanctioned the current study.
PICALM RNA expression in human post-mortem brains
Frozen human brain samples from the frontal cortex BA10 region were obtained via the NIH biobank from the Harvard Brain Tissue Resource Center and the University of Miami Brain Endowment Bank. For assessing PICALM expression, grey matter was carefully isolated from brain blocks. RNA was extracted using the Direct-zol RNA MiniPrep Kit and reverse transcribed into cDNA using the High-Capacity cDNA Reverse Transcription Kit, following the manufacturer’s guidelines. Quantitative PCR reactions were prepared with PowerUp SYBR Green Master Mix, using the QuantStudio Real-Time PCR System. Data analysis utilized the 2^{-\Delta \Delta C_{t}} method and were normalized to COTL1. Primer sequences were specified for PICALM and COTL1.
PICALM staining of hMGs
Paraffin sections (6 µm) from post-mortem human brain samples were accessed from UCLA or RUSH Alzheimer’s Disease Center repositories. The RADC samples were part of the Religious Orders Study and Rush Memory and Aging Project, with all RADC participants consenting to clinical evaluations and brain donations at death. Both studies received approval from the Rush University Medical Center IRB. The sections underwent a series of processing steps including incubation at 60 °C for deparaffinization and rehydration. Epitope retrieval utilized buffer at 95 °C, followed by treatment with sodium borohydride. Various blocking procedures and antibody staining methods were carried out, culminating in slides being mounted and images captured via an automated microscope.
iMG differentiation from human iPS cells
iMGs were generated from human iPS cells following previously described methods. Once the cells reached about 80% confluency post-passaging, they were dissociated and plated for differentiation. Various media were prepared for both embryoid body and iMG production, with continuous culturing leading to the creation of primitive macrophage progenitors.
iAst cell differentiation from NPCs
Neural progenitor cells derived from iPS cells were placed in astrocyte medium to generate astrocytes. Careful attention was given to cell density and dissociation to obtain a uniform astrocyte population, which involved multiple feeding and expansion processes.
Differentiation of glutamatergic neurons
A well-documented protocol was followed to differentiate iPS cells into glutamatergic neurons. Post-dissociation, the cells were transfected with lentivirus cocktails, leading to significant neuronal development over a span of days.
Differentiation of dopaminergic neurons
This crucial differentiation stage involved a carefully timed introduction of dopaminergic priming medium, leading to successful neuron maturation after a month.
Differentiation of GABAergic neurons
GABAergic neurons were developed from NPCs, employing a specific viral cocktail to enhance differentiation. The process required multiple transitions in medium and treatment, ensuring the generation of mature neurons.
Immunofluorescence staining of iMG and iAst cells
To study iMG and iAst cells, fixed samples were subjected to a series of staining and imaging protocols. Various primary and secondary antibodies were tested, and samples were analyzed through fluorescence microscopy.
RNA isolation and sequencing
RNA extraction from iPS, iAst, and iMG cultures took place using the RNeasy Plus Kit, followed by cDNA synthesis. RNA-seq was conducted on an advanced sequencing platform with specific targets for sample reads.
RNA-seq data and differential expression analyses
RNA-seq data underwent rigorous processing, involving alignment against the human genome and batch effect corrections. Statistical analyses determined gene expression changes across diverse samples.
ATAC-seq
Sample preparation for ATAC-seq was carried out following acknowledged methodologies, with conditions set for ideal cell viability and DNA extraction.
ATAC-seq data analysis and peak calling
A comprehensive data analysis of ATAC-seq results was conducted, utilizing specialized software for effective mapping and peak identification, alongside removing misaligned reads.
ASoC mapping
ASoC analysis aimed to identify functional variants based on chromatin accessibility differences, leveraging advanced genotyping techniques to rigorously categorize SNP variability.
sLDSC analysis of GWAS enrichment
Stratified linkage disequilibrium score regression was used to understand the genetic relationships in relation to various psychiatric and non-psychiatric disorders.
Torus GWAS enrichment analysis
A Bayesian hierarchical model performed SNP-based enrichment assessments, determining whether ASoC SNPs held significant relevance across different diseases.
CRISPR–Cas9 editing of human iPS cells
CRISPR guide RNA sequences were meticulously designed for specificity, followed by extensive cloning and transfection efforts to ensure successful gene editing in iPS cells.
Quality control of the CRISPR-edited iPS cell lines
Numerous strategies, including eSNP-karyotyping and specific antibody staining, were employed to verify the integrity and pluripotency of edited cell lines.
CRISPRoff epigenome editing of human iPS cells
Using CRISPRoff technology, researchers repressed PICALM expression in iMGs, assigning specific gRNA sequences for targeted epigenetic alterations.
CRISPRa to overexpress PICALM
Through the CRISPR-ERA method, gRNAs for PICALM activation were developed and tested for efficiency to facilitate the generation of models for overexpression studies.
Gene expression analysis by qPCR
qPCR procedures involved precise reverse transcription followed by robust amplification processes to characterize gene expression levels across various cell types.
Myelin isolation from mouse brains for phagocytosis
Myelin extraction followed a detailed protocol involving homogenization and centrifugation steps to isolate myelin effectively.
Phagocytosis assay for iMGs
iMG cultures underwent a series of tests to evaluate their phagocytic capabilities using specific labelled reagents for visual confirmation through microscopy.
ChIP
Chromatin immunoprecipitation processes were adapted from existing protocols to facilitate detailed analyses of protein-DNA interactions.
Fatty acid (Red-C12) transfer assay
Cells were prepared and incubated with specifically labelled fatty acids to assess lipid transfer and interaction under controlled conditions.
LD staining with BODIPY for iMGs
Staining procedures for iMGs involved fixation and subsequent dye application to evaluate lipid composition quantitatively.
LD staining with LipidTOX for iMGs
Similar methodologies as above were employed for staining and quantification of lipid droplets using LipidTOX technology.
ROS staining
iMG cultures underwent various treatments to assess reactive oxygen species levels, involving incubation with specific fluorescent dyes.
Lipid peroxidation assay using BODIPY C11
This assay measured oxidative stress levels in cultured iMGs, utilizing capable fluorescent indicators to assess lipid peroxidation status.
Filipin staining for iMGs
Filipin staining processes highlighted cholesterol levels in iMGs through specific fluorescent measures, followed by detailed imaging analyses.
Lysosomal staining for iMGs
Lysosomal markers were used to visualize and analyze lysosome functionalities in iMGs through precise dye applications followed by imaging.
FACS sorting of iMGs
FACS was utilized to isolate specific iMG populations based on surface markers, ensuring thorough quantitative evaluations post-analysis.
PICALM KO by CRISPR–Cas9 editing C20 cells
C20 cells were maintained and subjected to gene-targeting strategies utilizing CRISPR-Cas9 to achieve specific genetic modifications.
Immunoblots for iMG and C20 cells
Protein analyses were conducted using immunoblotting techniques to assess target protein levels in various cell types.
Myelin phagocytosis for C20 cells
Assays were performed to evaluate myelin uptake in C20 cells using specific markers to track phagocytosis visually.
LD staining for C20 cells
Live imaging of C20 cells was performed to document lipid droplet dynamics utilizing fluorescent markers.
Organelle marker staining for C20 cells
C20 cells underwent a series of staining protocols directed at visualizing organelles, with subsequent imaging for detailed analysis.
Lipid extraction from iMGs
Lipid extraction involved careful processing of iMG cultures, following well-established methods to obtain accurate lipid profiles.
Lipidomics analysis of iMGs
A comprehensive analysis of lipid profiles was conducted utilizing state-of-the-art mass spectrometry techniques, providing insights into lipid compositions.
Imaging quantification and statistical analyses
Quantitative analyses of imaging data were performed, employing rigorous statistical methods to ensure the systematic evaluation of findings.
Reporting summary
For detailed information regarding research design, refer to the linked Reporting Summary.