Remote recruitment of participants through the GHS platform
The study protocol received approval from Advarra (IRB no. Pro00074093). Participants were recruited via Google Health Services (GHS) and the Fitbit applications. GHS serves as a platform for conducting digital studies, allowing participants to enroll, check their eligibility, and provide informed consent. It also enables data collection from devices like Fitbits and Pixel watches and allows users to complete questionnaires and arrange blood tests through Quest Diagnostics. As part of the consent process, participants were required to sign the Quest HIPAA authorization within the GHS app. This study enrolled 4,416 participants in the USA, with a subgroup of 1,165 included in the analysis due to complete data availability. The study was mainly conducted remotely, requiring participants to visit a Quest Patient Service Center only for blood draws. Participants were asked to wear their devices continuously, schedule blood draws, and respond to questionnaires.
Inclusion criteria included: participants aged 21 to 80 residing in the USA; Android Fitbit users with a tracking device; users needing at least three months of data with 75% usage to monitor activity and sleep; those willing to update their Fitbit app; those prepared to link their Quest Diagnostics account with the GHS app; participants able to understand and communicate in English; and those with access to a Quest Diagnostics location.
Exclusion criteria involved participants from Alaska, Arizona, Hawaii, and certain US territories, as well as those with uncontrolled medical conditions, or issues that could complicate blood sample collection.
Study design
Participants linked their Fitbit accounts to the GHS app and allowed GHS to collect Fitbit data throughout the study, including data up to three months prior to enrollment. After enrolling, participants were required to: (1) continuously wear their devices for a minimum of three out of four days; (2) complete four questionnaires addressing demographic details, health history, personal health perceptions, and blood test interpretations; (3) make an appointment for blood test orders within 65 days of enrollment; (4) visit a Quest Patient Service Center for a blood draw; and (5) review their lab results via the GHS app when available.
Collection of metadata
Demographic information (like age, gender, weight, and height) along with optional measures such as medical conditions and self-reported health management, were collected through a questionnaire completed by participants promptly after enrollment via the GHS app.
Blood biomarker measurements
Eligible participants scheduled visits to any nearby Quest Diagnostics Patient Service Center for a standard blood draw. The tests measured various health indicators such as complete blood count and cholesterol levels. Every participant was required to fast for at least eight hours prior to the draw, ideally in the morning from 7:00 to 10:00 AM, to reduce variability due to the diurnal cycle. The study had physician oversight, with lab results returned to participants through GHS and not stored there. Results remained available in the participants’ Quest Accounts post-study.
Wearable data collection
Participants were asked to wear their own Fitbits or Pixel watches, sharing a variety of data, including heart rate, physical activity metrics, sleep patterns, skin temperature, and blood oxygen levels. They consented to share this data regularly, allowing for a comprehensive understanding of health metrics related to their activities and sleep.
Selection of HOMA-IR thresholds
The criteria for HOMA-IR thresholds were informed by previous studies suggesting values between 1.5 and 3 for insulin resistance, with a selected threshold of 2.9 falling in the middle of the ranges studied. Those displaying HOMA-IR values between 1.5 and 2.9 were classified as insulin-resistant.
Modeling and computational pipeline
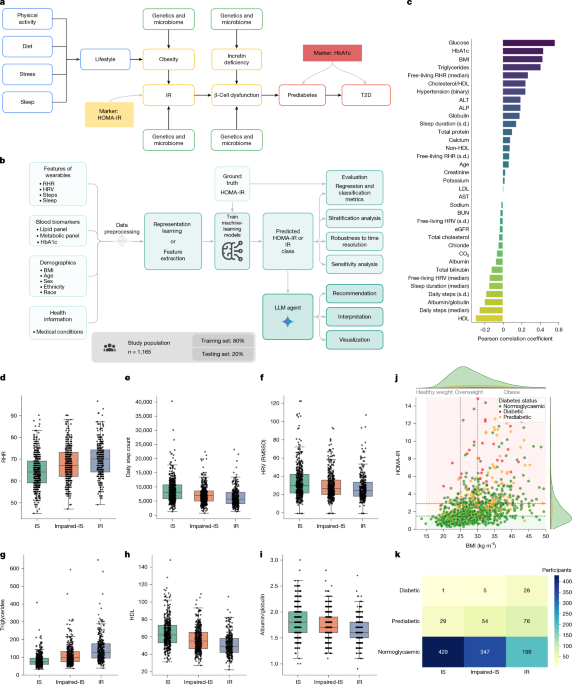
Our approach consists of four phases: (i) preprocessing the data; (ii) modeling and training; (iii) predicting and classifying HOMA-IR; and (iv) interpreting results via large language models. Below, we elaborate on each of these phases, incorporating strategies for evaluation and extensive ablation studies.
Data preprocessing
Demographics
Data from user surveys allowed us to extract users’ ages and calculate BMI based on their height and weight. We excluded users with a BMI beyond acceptable limits for quality assurance.
Digital markers
We aggregated digital markers from Fitbit data over specified time frames pre-blood test, exploring the optimal consolidation period through ablation studies.
Blood biomarker data
For participants not fasting as specified or with missing data, we excluded them from analysis. We also eliminated any outliers based on the true calculated HOMA-IR values.
Data standardization
The dataset for modeling combined demographics, digital markers, and blood biomarker information. For consistency, we standardized input features to achieve zero mean and unit variance across training and testing subsets.
Modeling
Our primary objective was to predict HOMA-IR values and classify individuals based on established thresholds, using regression techniques which facilitate deeper analysis and interpretation of individual data.
Direct regression
We utilized gradient boosting machines, specifically the XGBoost framework, which is adept at managing complex datasets. XGBoost offers efficiency and scalability, enabling us to leverage both linear and non-linear learning approaches based on our feature interactions.
Representation learning
While direct regression can handle dimensional data, we assessed whether more concise data representations could enhance predictive performance. Relevant literature supports the importance of well-structured health data for various applications.
MAE
We employed masked autoencoders (MAEs) for representation learning because they have demonstrated improvements over previous methods in recreating data formats through reconstruction objectives.
Model training and HOMA-IR predictions
Regression results stemmed from two stages: representation learning followed by training an XGBoost model using learned representations to predict HOMA-IR values. We trained both models across specified epochs, utilizing the Adam optimizer and incorporating adaptations based on performance metrics.
Parameter selection
To identify optimal parameter sets for our models, we underwent a grid search, performing five-fold cross-validation to evaluate each combination’s effectiveness.
Independent validation cohort
This study’s protocol, approved by WCG (IRB no. 1371945), involved a broader longitudinal study assessing lifestyle impacts on health metrics. Collected data included various anthropometric measurements and biomarker analyses. Participants attended initial and concluding appointments for assessments and were asked to wear Fitbit devices consistently.
Inclusion criteria specified California residency, age, ability to stand and walk unaided, informed consent willingness, smartphone ownership, and device comfort. Exclusion factors involved uncontrolled health conditions, impracticalities regarding blood samples, and several other medical and situational criteria.
Of the 144 initially enrolled individuals, 72 provided complete data and served as the validation cohort. We ensured that their wearable data adequately represented their lifestyle choices during the final assessment period.
Evaluation of prediction and classification of HOMA-IR
Regression evaluation
To assess HOMA-IR prediction performance, we evaluated the continuous values, calculating average mean absolute error across test folds to analyze prediction consistency.
Classification evaluation
Insulin resistance classification was based on continuous HOMA-IR values, using a threshold of 2.9. We reported various metrics across test folds to gauge effectiveness, including sensitivity and specificity.
Time-dependent sensitivity analysis
To understand predictions’ robustness over different time frames, we implemented rolling window aggregation over multiple days, reporting prediction stability across the periods we examined.
IR agent
Our large language model (LLM) agent, aimed at improving human-computer interactions within health contexts, incorporates multi-step reasoning and planning capabilities relevant to interpreting health metrics.
This agent dynamically processes incoming queries by leveraging pertinent information and tools, adapting to real-world complexity and ensuring user engagement through relevant responses.
IR agent toolbox
Our IR agent utilizes a variety of tools for effective query resolution, including web searches and specialized models for precise computations, significantly reducing error potential and improving response reliability.
Implementation of the IR agent
Implemented via Google DeepMind’s OneTwo library, our IR agent’s structure supports interactions and reasoning required to continuously refine health-related responses.
Evaluation of the IR agent
We aimed to measure the advantages of including HOMA-IR predictions in our agent’s responses against standard LLM outputs, focusing on accuracy and depth as evaluated by medical experts.
Statistical analysis
Statistical rigor was maintained through established testing methods and Python for model training and evaluation.
Visualization methods
We employed various visualization tools for data representation throughout the project.
Reporting summary
Further information regarding the research design can be found in the linked summary.