Mice
All experimental procedures followed the ethical guidelines set by the Institutional Animal Care Committees at the University of California, San Francisco (UCSF) and the South Australian Health and Medical Research Institute (SAHMRI), adhering to the NIH and NHMRC standards regarding laboratory animal care. Both male and female mice, aged between 8 and 16 weeks, were utilized and given unrestricted access to standard lab chow and sterile water. They were kept in a controlled setting with a 12-hour light-dark cycle. TRPM5-GFP mice were employed for the visualization of tuft cells (a gift from R. F. Margolskee). Furthermore, Pou2f3−/− mice from Jackson Laboratory (strain 037040) facilitated tuft cell knockout. A conditional knockout of Chat in intestinal tuft cells was achieved by crossing VilCre mice (strain 021504 from Jackson Laboratory) with Chatflox mice (provided by J. Chan). To perform gGRAB5-HT3.0 sensor imaging, VilCre mice were crossed with the Rosa26gGRAB-5-HT3.0-P2A-jRGECO1a reporter line. For GCaMP imaging in organoids, Tac1Cre mice (strain 021877 from Jackson Laboratory) were crossed with Polr2aGCaMP5g-IRES-tdTomato mice (also provided by L. Jan, Jackson Laboratory, strain 024477). To inhibit serotonin release from enterochromaffin (EC) cells, the tetanus toxin was expressed in these cells using the Cre and Flp-dependent mouse model RC::PFTox (gift from S. Dymecki). Specifically, ePetFlp hemizygous/RC::PFTox homozygous mice were crossed with Tac1Cre homozygous mice to create Tac1Cre;ePetFlp;PFTox mice. For nerve fiber recordings, Scn10aCre mice (gift from W. Imlach, Jackson Laboratory, strain 036564) were crossed with the ChR2 reporter line (Rosa26lsl-ChR2; Jackson Laboratory, strain 012569). Genetically modified and control mice (either littermates or age-matched) were randomly selected for all behavioral testing.
Crypt cell isolation and organoid culture
Adult male Tac1Cre;Polr2aGCaMP5g-IRES-tdTomato, TRPM5-GFP, and Pou2f3−/− mice were employed to generate intestinal organoids, as previously described. Approximately 8-cm segments of the ileum were used to establish TRPM5-GFP organoids. To avoid ectopic expression of Tac1Cre in the lower intestine, the upper jejunum was used for Tac1Cre;Polr2aGCaMP5g-IRES-tdTomato organoids. These organoids were maintained and passaged every six days in organoid growth medium (advanced DMEM/F12 supplemented with penicillin-streptomycin, 10 mM HEPES, GlutaMAX, B27, 1 mM N-acetylcysteine, 50 ng/ml mouse recombinant epidermal growth factor, R-spondin 1, and mouse Noggin).
Cell lines
The R-spondin-1-expressing HEK293FT cells were kept in DMEM, 20% fetal calf serum, 1% penicillin-streptomycin, and 125 µg/ml zeocin at 37 °C, 5% CO2. After producing R-spondin 1 conditioned medium, zeocin was removed. HEK293T cells were cultured in DMEM, 10% FCS, and 1% penicillin-streptomycin at similar conditions and were transfected with Lipofectamine 3000. For biosensor experiments, 200 ng pDisplay-gGRABACh4h-IRES-mCherryCAAX or 200 ng pcDNA3-hM1R-P2A-GCaMP8m was transfected into HEK293FT cells in 24-well plates. For the 5-HT3 biosensor experiment, 200 ng pcDNA3-5-HT3A and 20 ng pcDNA3-mApple were co-transfected into HEK293FT cells in 24-well plates.
Induction of tuft cell hyperplasia
Organoid tuft cell hyperplasia was induced by exposing organoids to 20 ng/ml IL-4 in the growth medium for two days, followed by one day in IL-4-free medium. The biosensor experiments took place on day six. To induce tuft cell hyperplasia in mice, they received intraperitoneal injections of 500 ng IL-25 on consecutive days. Tissues for imagingwere collected on day five.
GCaMP imaging using intestinal organoids
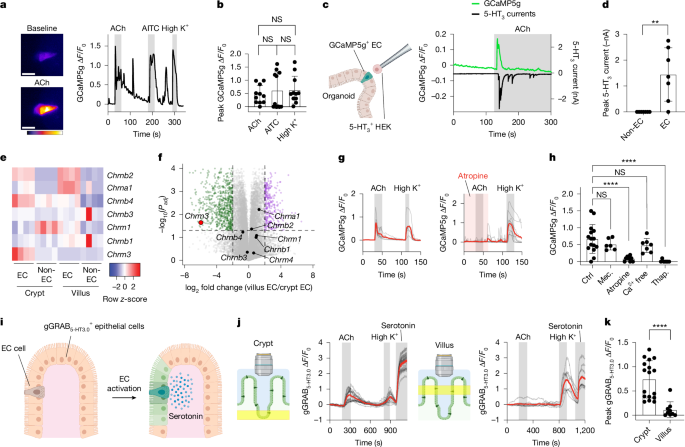
Five days post-passage, Tac1Cre;Polr2aGCaMP5g-IRES-tdTomato organoids were separated from Matrigel and mechanically disrupted. The fragments were plated onto Cell-Tak-coated coverslips and placed in a recording chamber filled with Ringer’s solution. EC cells were identified through tdTomato expression. GCaMP imaging utilized an upright microscope with a Grasshopper 3 camera and was maintained under continuous Ringer’s solution flow. All pharmacological reagents were administered via local perfusion. The acquired images underwent analysis using Fiji.
M1R–GCaMP8m biosensor experiments
Following the tuft cell hyperplasia induction protocol described earlier, each well in a 24-well plate had the organoid medium replaced with Ringer’s solution to remove residual growth medium. After a brief incubation period, the fresh Ringer’s solution was replaced and allowed to sit for a further two hours before imaging. For M1R–GCaMP8m biosensor cell preparation, HEK293FT cells that were transfected with pcDNA3-M1R-P2A-GCaMP8m were isolated and plated on glass coverslips. Imaging occurred using a similar setup. The entire surface area of each biosensor cell was utilized for calculating ∆F/F0 values.
gGRABACh4h biosensor experiments
HEK293FT cells transiently transfected with pDisplay-gGRABACh4h-IRES-mCherryCAAX underwent dissociation, washed, and plated atop intestinal organoids. For isolated tuft cell and TRPM5-GFP organoids biosensor experiments, individual HEK293FT cells were carefully moved close to tuft cells using a glass pipette. Imaging was executed using the same setup, focusing on the entire biosensor cell area for ∆F/F0 calculations. The bath solution remained static to avoid washout of endogenously released ACh.
5-HT3 biosensor experiments
HEK293FT cells transfected with pcDNA3-5-HT3A and pcDNA3-mApple were isolated, washed, and plated on intestinal organoids. The transfected cells were identified through mApple expression, and imaging took place while maintaining a whole-cell configuration. During the imaging, the potential was held at −80 mV, with careful placement next to EC cells using a glass pipette. Pharmacological agents were manually applied, and all images were properly analyzed using Fiji.
Patch-clamp recordings
Patch-clamp recordings utilized pClamp software with a Multiclamp 700A amplifier. Electrodes were created from borosilicate capillary and the intracellular solution comprised mainly of K-aspartate, NaCl, and other components. The objective was to measure voltage-gated calcium currents, utilizing specialized external and internal solutions. Agonists were delivered via gravity-driven microperfusion.
Ex vivo serotonin sensor imaging
Segments of the jejunum were isolated from genetically modified mice for imaging. Tissue was prepared for stability before imaging with a confocal microscope. The setup included a perfusion system for buffering solutions and pharmacological agents. Measurements involved activating the gGRAB5-HT3.0 sensor with serotonin and evaluating baseline serotonin levels after infection.
Dissociation of tuft cells
The ileum from the TRPM5-GFP mice was segmented and treated for cell dissociation. The epithelial layer was subjected to digestion, washed through several stages, and the cells were kept on ice for further usage.
Dissociation of EC cells
To isolate EC cells for GCaMP imaging, intestinal tissue was prepared, and the villus epithelium was dissociated using specific methods. The remaining epithelium underwent subsequent treatments before the cells were filtered and prepared for imaging.
Brainstem tissue collection
Mice underwent treatment with IL-25 before being euthanized for tissue collection. They were perfused with saline and paraformaldehyde before their brains and brainstems were isolated for further analysis.
Histology and immunostaining
Immunofluorescence was conducted on tissue sections, including organoid fixation and staining with various antibodies. Images were captured using advanced microscopy techniques, with detailed attention paid to capturing high-resolution images.
In situ hybridization
Cryosections were prepared for RNA fluorescence in situ hybridization (RNA-FISH) using specific probes. Imaging took place with appropriately set confocal microscopy, and maximum-intensity projections were analyzed.
Brainstem FOS quantification
The medial part of gut-innervating vagal afferents was selected for quantifying activated neurons. Carefully recorded data points represented FOS+ neurons averaged across multiple slides, ensuring a representative analysis.
Food intake assay
Mice were acclimated to metabolic cages for measurements of food intake alongside control groups. After establishing baseline data, injections with IL-25 or saline were carried out to observe changes in food consumption.
For acute tuft cell stimulation, mice were housed individually and acclimated to automated feeding devices. Following this, an experimental protocol monitored food intake after treatment with specific agents.
During Nb infections, mice underwent similar adjustments with feeding devices to measure their consumption patterns post-infection.
Ex vivo nerve fiber recordings of mucosal sensory afferents innervating the small intestine
Recordings in the jejunum involved careful dissection and preparation, ensuring the neurovascular bundles were properly positioned for recording. Action potentials were analyzed after being captured through designated equipment.
Optogenetic stimulation of jejunal mucosal afferents
Mice with specific genetic modifications allowed for precise optogenetic activation of mucosal afferents for studying responses when stimulated by light. Drugs were perfused sequentially to assess their impact on the generated action potentials.
Behavioural assays
Behavioural assessments were conducted on mice after inducing tuft cell hyperplasia through injections over several consecutive days. Mice were tested for various locomotor activities and behaviors in a specialized environment with monitoring technology.
Abdominal electronic von Frey Hair testing
To evaluate mechanical allodynia, the electronic Von Frey hairs were employed with mice subjected to treatment. Testing allowed for measuring abdominal withdrawal thresholds following specified protocols.
Assessment of nest building
Nesting behaviour was observed in mice subjected to IL-25 treatments, with specific nests assessed according to established criteria. The results helped gauge animal well-being, considering their natural instinct to nest.
Retrograde tracing of mucosal afferents from the small intestine
For tracing vagal ganglia collection, adult male mice underwent specific conditions including retrograde tracing via cholera toxin. This enabled targeted investigations of neuronal responses in subsequent analyses.
Cell culture for calcium imaging and single-cell RT–PCR of small-intestine mucosal-traced vagal neurons
Vagal ganglia were isolated and cultured in specialized conditions post-tracing, aimed at facilitating calcium imaging and single-cell transcription analysis.
Single-cell RT–PCR of mucosal-traced vagal ganglion neurons
Traced neurons were manually selected for further genomic analysis following specified protocols, with controls included for accuracy.
Calcium imaging of small-intestine mucosa-innervating vagal ganglia neurons
Post-isolation, neurons were treated for imaging following careful preparations, enabling the assessment of calcium responses in reaction to various stimuli.
Analysis of single-cell and single-nucleus RNA-sequencing datasets
Processed RNA-sequencing data for specific mouse models were accessed, focusing on well-annotated cellular subtypes, enabling in-depth expression profiling.
Statistical analysis
Data assessment utilized systems ensuring reliability, with careful consideration of parameters for determining significant results.
Reporting summary
For comprehensive details on research design, refer to the linked Nature Portfolio Reporting Summary.