Ethics statement
This study involving human sera received approval from the Scientific and Ethics Review Unit of the Kenya Medical Research Institute (protocol SSC 3426). Donors provided individual consent for their blood samples to be used in research.
Construction of gene libraries
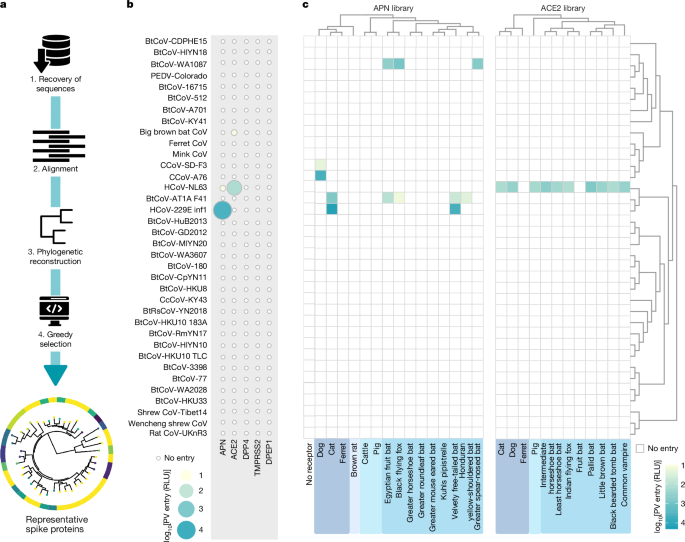
Figure 1a outlines the analysis pipeline employed for extracting alphacoronavirus data and constructing the spike protein library utilized here (n = 40). We gathered all publicly accessible alphacoronavirus genome sequences from the Virus Pathogen Database and Analysis Resource hosted by the Bioinformatics Resource Center at the National Institute of Allergy and Infectious Diseases. As of May 2021, this database comprises 19,082 alphacoronavirus genomes, allowing us to extract sequences from the entire spike protein-coding region, ultimately producing a dataset of 2,714 sequences. The spike protein-coding DNA sequence alignment was formed using MAFFT (v.7.526), combining structural alignments from homologous spike protein structures sourced from the UniProt Reference Clusters. Phylogenetic reconstruction was accomplished via IQTREE (v.2.3.4), with 1,000 ultrafast bootstrap replicates (UFBoot) and 1,000 SH-like approximate likelihood ratio tests. Codon model reconstruction included selecting the best-fit model using IQTREE’s model selection process. Patristic distances were computed in R (v.4.4.1) using the ape package, aiding the selection of the n = 40 spike protein-coding sequences through a greedy algorithm. In essence, the greedy algorithm maximized both minimum phylogenetic distance and phylogenetic diversity based on the assumption of evolutionary units. We noted Faith’s phylogenetic diversity of the induced minimal subtree and compared it against 10,000 random panels of equivalent size. Interestingly, a receptor-binding domain (RBD) was challenging to identify for two unclassified viruses, as well as those categorized in the Soracivirus and Luchacovirus subgenera.
Plasmids used for pseudotyping
Codon-optimized synthetic genes for selected alphacoronavirus spike protein-coding sequences were ordered from Biobasic, subcloned in pcDNA3.1, and tagged with HA at the C-terminus. The APN library genes came from GenScript and were subcloned with a N-terminal V5 tag in pCAGGS. For ACE2 library and human DPP4, the ectodomains of the ORFs were tagged with a N-terminal HA in pDisplay. The CEACAM libraries were obtained from GenScript and subcloned into pcDNA3.1(+) with a C-terminal 8×His tag. We also cloned Flag-tagged human TMPRSS2 and human DPEP1 into pcDNA3.1. The GenBank accession numbers for the sequences used in this study can be found in various supplementary tables.
Cells
Mycoplasma-free HEK293T (human kidney), Calu3 (human lung), Caco2 (human colorectal adenocarcinoma), and HuH7 (human hepatoma) cells were cultured in DMEM with 10% FBS, penicillin, streptomycin, and sodium pyruvate. Additionally, mycoplasma-free THP1 (human monocytes) and LCL (human B lymphocytes) cell lines were sustained in suspension using RPMI1640 with 10% FBS. All reagents for culturing were sourced from Gibco, and all cells were maintained in a humidified environment at 37 °C with 5% CO2.
Pseudotype virus production
To pseudotype alphacoronavirus spike proteins, plasmids encoding their ORF were transfected into HEK293T cells using polyethylenimine (PEI). We included HIV-1-based lentiviral vectors containing structural proteins and a luciferase reporter gene during transfection. On the following day, we replaced the medium and collected pseudoviruses from the supernatant 48 and 72 hours post-transfection, pooling them together. Centrifugation was then employed to eliminate cellular debris, and pseudoviruses were stored at −80 °C. Purification of pseudoparticles was conducted through ultracentrifugation, and spike protein expression was analyzed by SDS-PAGE. Notably, while some spike proteins demonstrated entry capability, the correlation between protein expression in the immunoblots and entry signals was not consistent. In cases where six spike proteins failed to pseudotype, we replaced them with others displaying at least 95% amino acid similarity. The only match was for PEDV-KDJ, substituted with PEDV-Colorado. The HCoV-229E spike protein sequence produced low levels of pseudotyping and was not effective for later applications, necessitating its replacement with a different strain. Original immunoblots are stored in the Zenodo repository.
Pseudovirus entry assay
For receptor usage tests, HEK293T cells were transfected with relevant receptor plasmids. After a day, we replaced the medium with fresh DMEM enriched with FBS, then dispensed cell preparation and pseudovirus preparation into 96-well plates. Two days later, the supernatant was removed, and cells were treated with Bright-Glo for luciferase signal acquisition. Additionally, to test cell line permissivity to pseudovirus, transduction occurred in confluent human cell lines with undiluted pseudoparticle supernatant. The assay to inhibit CEACAM6-mediated entry involved pre-incubating pseudotyped CcCoV-KY43 spike protein with specific monoclonal antibodies or recombinant human CEACAM6 before introducing HEK293T cells transfected with pcDNA-human CEACAM6.
Flow cytometry for expression of the receptor libraries
Plasmids were transfected into HEK293T cells using Trans-IT X2. The ensuing day, cells were fixed and permeabilized. After washing, they were incubated with specific anti-tag antibodies for 1 hour prior to flow cytometry analysis. To evaluate the specificity of certain antibodies, HEK293T cells were transfected with a diverse library of human CEACAM proteins, followed by washing, fixation, and antibody incubation involving anti-human CEACAM6 monoclonal antibodies. Gating strategies were defined for data analysis.
Human virus receptor discovery
Human receptor ectodomains were expressed as biotinylated proteins through co-transfection in HEK293 cells. After five days of expression, supernatants containing the proteins were collected, and heparin and imidazole concentrations were adjusted for purification at 96-well HisTrapHP plates. The procedure included various washing and elution steps to ensure purity. The assessments of protein concentrations and purity followed standard methods, leading to several experiments targeting interactions between CEACAM proteins and the viral RBDs.
Site-directed mutagenesis
Substitutions in CEACAMs were performed using a specific kit, with primer designs facilitated through an online tool.
Cell–cell fusion assay
HEK293T cells were transfected with constructs for human CEACAM receptors or spike proteins and were then co-cultured in specified ratios. After maintaining them for two days, we lysed the cells and quantified luciferase signals to measure membrane fusion.
Knockdown of CEACAM6 in human cells
CEACAM6-targeting siRNA was introduced in specific cells through electroporation, followed by pseudotyped spike protein introduction. Expression reduction was confirmed with Western blots, and stable cell lines with knocked down CEACAM6 were also generated, utilizing established protocols for shRNA and subsequent selection processes. The original immunoblots are available in the Zenodo repository.
Transcriptomic analysis of available datasets
We downloaded single-cell RNA-sequencing data from the Human Protein Atlas, covering 31 human organs. The matrices were processed using Seurat in R, also incorporating data gathered from the Human Lung Cell Atlas for targeted lung-specific analyses.
Recombinant protein production
We constructed various CEACAM constructs and full-length spike proteins in appropriate vectors. Expi293 cells were cultivated following specified conditions, and the necessary steps for protein purification were executed, ensuring assessments of the targeted proteins’ quality and presence.
Protein binding determined by ELISA
ELISA procedures were employed to determine binding affinities, with various incubation and washing stages outlined. Binding assessments were conducted with human sera, alongside statistical analyses depicting relationships between antibody responses.
Flow cytometry for receptor–viral protein binding
To explore viral protein binding, we transiently transfected HEK293T cells with various receptor plasmids and then assessed how effectively they bound to certain viral proteins, continuing with detailed data analysis techniques.
Crystallization
Complexes of CEACAM6 with specific RBDs were formed and subsequently crystallized in a nanolitre-scale format, leading to further preservation steps before data collection.
X-ray data collection and structure determination
Notably, we collected diffraction data using various beamlines, applying precision integration and scaling methods for accurate results. Both complexes showed high affinities and purities, supporting substantial analysis and interpretation of structural data.
Biolayer interferometry
This technique allowed us to compare binding affinities of CEACAM proteins for the examined RBDs, employing sophisticated methodologies to ensure reliable results.
ITC
Measurements of binding affinities were performed using an advanced calorimeter, yielding significant insights into protein interactions.
C. cor and human population distribution data in Kenya
The GPS data from bat survey locations in Kenya were systematically arranged and classified, facilitating informative analyses regarding species distributions.
Phylogenetic reconstruction
Spike gene sequences were aligned and calibrated effectively, following a detailed Bayesian analysis methodology that captured the evolutionary history of alphacoronaviruses and CEACAM6 usage.
Additional computational analysis
Various molecular graphics and statistical functions were applied to analyze results thoroughly, ensuring valid interpretations of pseudotype entry effects and overall study findings.
Biosecurity statement
Acknowledging the health risks linked to CEACAM6 identification, we took steps to ensure preparedness and enhanced biosecurity measures in Kenya, emphasizing the ethical considerations at every stage of our research.
Reporting summary
Further design information related to this research can be found in the linked Nature Portfolio Reporting Summary.