Experimental methods
Human sample preparation
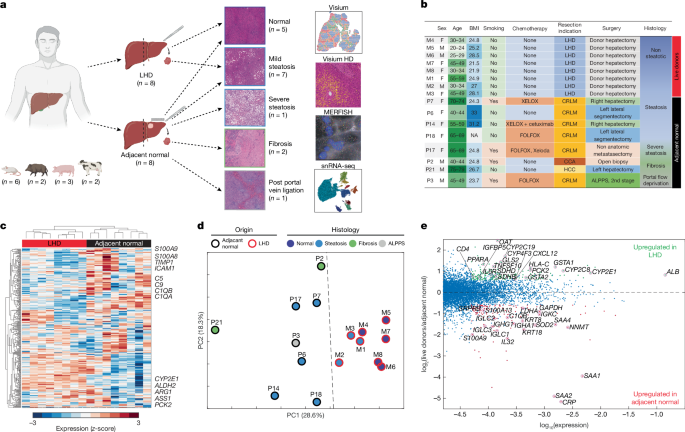
Human samples and clinical data were collected from patients who underwent either hepatectomy for liver conditions or donor hepatectomy for living liver donation. In surgeries related to liver pathologies, samples were taken at least 5 cm away from the pathological margin. For donor surgeries, liver biopsies of 1 cm³ were acquired right after the abdominal cavity was opened, before the liver was moved or surgery started. These procedures took place at Sheba Medical Center in Israel and the Mayo Clinic in Rochester, MN, following ethical approvals from their respective committees. Informed consent was acquired from all patients, and all practices adhered to Helsinki guidelines. Samples were only collected if no issues were found in the sampling area during evaluations. Approximately 1 cm of liver tissue, away from the capsule, was removed. The tissue samples were gently washed in PBS and either embedded in OCT for Visium and MERFISH analysis or frozen for Visium HD analysis.
Non-human sample preparation
Whole livers from pigs were procured from a butcher in Kibbutz Lahav in Israel, which utilizes surplus animals. These included adult male and female pigs, both about 5.5 months old. Liver sections from cattle were donated by a butcher in Haifa, and included adult males around 17 months old. Additionally, liver samples from wild boars were sourced from animals euthanized through regulated population control efforts by the Israel Nature and Parks Authority (INPA). A permit was obtained prior to this process, and the researchers did not influence the circumstances of the animals’ death. Ages and weights of the boars were estimated by experienced rangers and a veterinarian. The tissues were similarly cleaned in PBS and prepared for spatial transcriptomics analysis.
10X Visium spatial transcriptomics
Fresh-frozen OCT samples were cryosectioned into slices 10 µm thick and placed onto Visium Spatial Gene Expression slides. Each slide had slices from four different patients, leading to a total of four slides for 16 human samples. These slides were fixed with chilled methanol, stained as per guidelines, and then imaged using a specific Leica microscope setup. The permeabilization process took about 24 minutes for all samples. After capturing the transcripts, the libraries were prepared and sequenced using an Illumina platform. The initial data was processed with specific software from 10X Genomics.
10X Visium HD spatial transcriptomics
Fresh-frozen tissue samples were converted into FFPE blocks, and 5 µm sections were mounted onto Visium HD slides for deparaffinization. H&E staining helped visualize tissue morphology, while high-res images were captured using a Leica microscope. Post-imaging, sections underwent procedures to destain and de-crosslink before preparing specific probes for targeting genes.
MERFISH spatial transcriptomics
Liver OCT blocks were cut into 10 µm sections and fixed with paraformaldehyde. Tissues underwent hybridization, gel embedding, and clearing guided by user protocols, utilizing a designated panel for pan-cancer pathways. After hybridization, samples were stained and prepared for imaging.
Immunofluorescence staining
Immunofluorescence was carried out on paraffin-embedded sections via a Leica detection platform. After sections were deparaffinized, antigen retrieval occurred before blocking non-specific binding. Primary antibodies against various proteins were then incubated, followed by exposure to secondary antibodies and visualization using a high-magnification imaging system.
PhenoCycler antibodies and imaging
The antibody panel used consisted of DAPI nuclear staining and 48 antibodies targeting various liver cell proteins. Validation of antibodies occurred on control tissues with available matched data. PhenoCycler runs were performed following the manufacturer’s instructions, ensuring optimal settings for signal detection.
Single-nucleus isolation from FFPE tissue samples
Three sections were washed with xylene to remove paraffin, followed by rehydration through consecutive ethanol washes. The sections were then treated to disrupt tissue and isolate nuclei using a specific digestion mix. Nuclei underwent filtering and centrifugation before being counted and stored appropriately.
Single-nucleus RNA profiling from FFPE tissue samples
Gene expression libraries were prepared in accordance with specific guidelines for nuclei obtained from FFPE samples. A hybridization procedure occurred, followed by washing and resuspension of nuclei, which were then prepared for sequencing using quality controls and calibrations.
Mouse immunofluorescence
Mouse experiments received institutional ethics approval and were carried out following relevant guidelines. Livers from two-month-old male mice were collected, fixed, and stored for later staining.
Computational methods
Genome alignment
Human and non-human samples were aligned to their respective genome assemblies for analysis.
Spatial transcriptomics preprocessing
To filter out outlier spots, criteria based on UMI counts and mitochondrial fractions were applied. Additional anatomical considerations were made during evaluations, ensuring appropriate filtering before analyses began.
Pseudo-bulk analyses of human samples
Specific gene sets were targeted to minimize batch effects, and expression levels were computed per patient. Various statistical methods were utilized for hierarchical clustering and PCA.
Human portal–central axis spot annotation
For Visium data, correlations between gene expressions were analyzed to create landmark gene sets for zonation scoring. These scores informed the classification and annotation processes.
Human hepatocyte marker gene identification
Markers for hepatocytes were identified based on existing single-cell data, focusing on normalized expressions and expression ratios between hepatocytes and other cell types. A comprehensive list of marker genes was generated.
Cross-species zonation reconstruction
Similar methodologies applied to human samples were adapted for non-human samples, facilitating comparative analyses of zonation profiles by genetic expression and weights.
Cross-species orthologous genes analysis
Data regarding orthologous genes across species were refined to allow for targeted comparisons, with a focus on identifying hepatocyte-specific gene expressions that were consistent between species.
Liver cell atlas using Visium HD
Visium HD processes involved single-nucleus segmentation and classification of cells based on morphological features. Establishing accurate classification and gene expression values followed detailed extraction protocols.
Liver cell atlas using snRNA-seq
snRNA-seq data from multiple patients were combined and analyzed to identify high-quality cells, with background subtraction performed to improve data reliability. Further analysis focused on gene expression patterns and zonation behaviors among cell types.
Ligand–receptor and transcription factor analysis
Computed correlations between ligands and receptors focused on specific pairs in the dataset. An analysis of transcription factors was conducted based on prior research references.
Lipid droplet quantification
The extraction of lipid content values from samples was done using classifier training on H&E images, focusing on identifying specific histological features within the data.
Comparison to mouse model of steato-hepatitis
Mouse data were analyzed for lobule layers, correlating expression levels between steatotic and non-steatotic conditions, using previously established methodologies and comparisons.
Ethics statement
All procedures were conducted following ethical standards, with approvals acquired from relevant committees and informed consent obtained from participants and caretakers of animal subjects.
Reporting summary
Additional details on the research design can be accessed via the linked reporting summary.