Background
The Democratic Republic of the Congo (DRC) is currently battling multiple health crises, including outbreaks of mpox, cholera, and malaria. On September 4, 2025, the DRC’s Ministry of Public Health officially announced its 16th Ebola Virus Disease (EVD) outbreak after confirming the presence of the Ebola virus (EBOV) in samples from a patient in the Bulape Health Zone of Kasai Province.
This marked resurgence of EBOV is not unexpected given that there have been 15 previous Ebola outbreaks in the country. In late August 2025, health authorities in Bulape raised alarms regarding cases of hemorrhagic fever, which resulted in fatalities among both patients and healthcare workers. Following these suspicions, specimens—including a buccal swab from a deceased patient and blood samples from five suspected cases—were sent to the national laboratory, Institut National de Recherche Biomédicale (INRB) in Kinshasa, for confirmation and genome sequencing.
Methods
On September 3, 2025, the INRB tested the six samples using the GeneXpert Ebola assay, the BioFire FilmArray System, and real-time PCR with the Altona RealStar Filovirus RT-PCR kit. Positive PCR samples were then sequenced using a nanopore sequencing device from Oxford Nanopore Technology.
In brief, viral RNA was extracted from 140 µL of inactivated samples following manufacturer guidelines using a viral RNA extraction kit from QIAGEN. Reverse transcription was conducted using LunaScript RT SuperMix from New England Biolabs. Multiplex PCR was carried out on cDNA using Q5 Hot Start High-Fidelity master mix and EBOV-specific primers designed for Mangina variants to amplify the EBOV genome. The preparation included a triplicate library using the rapid sequencing DNA V14 barcoding kit from Oxford Nanopore Technologies, and was subsequently sequenced on a GridION sequencer. Base calling was completed using the High accuracy model from Guppy, and the iVar tool was employed for trimming sequences and generating a consensus genome.
Results and Discussion
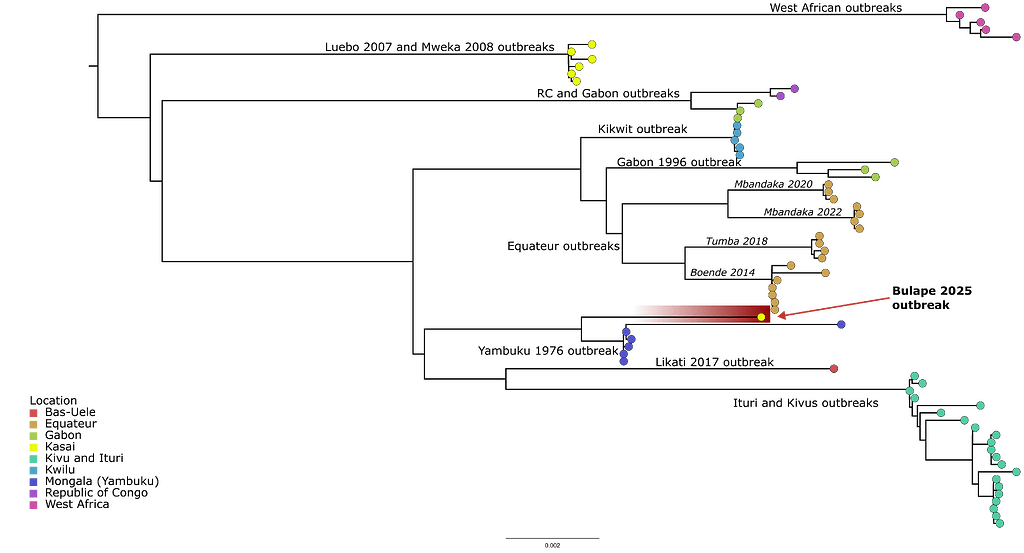
Within an hour of library preparation, we managed to produce an accurate consensus genome. The newly generated EBOV genome is 99.97% complete, comprising 18,819 nucleotides. We conducted a multiple sequence alignment of 71 complete EBOV genomes, including this new one, using MAFFT. A phylogenetic tree was then constructed using IQ-TREE v2.1.4. The new genome shows a nucleotide similarity of 99.52% to the genome from the Ebola virus identified in 1976, indicating that this case likely reflects a new zoonotic spillover and is not associated with the outbreaks in Luebo (2007) or Mweka (2008/2009).
Acknowledgements section
Our gratitude goes to the ARTIC Network and the University of Nebraska for supplying the necessary primers. Additionally, we appreciate Culmen International for providing the BioFire panels and primers through a cooperative agreement funded by the US CDC. Support from various organizations, including the Africa Pathogen Genomics Initiative, has been invaluable for genomic surveillance efforts in the DRC.
Statement on ongoing work and analysis prior to publication
This genome is being shared before publication, and it’s important to note that this data is preliminary; our analyses are still ongoing, and we are preparing a publication. For those interested in using our data pre-publication, please reach out to Prof. Placide Mbala-Kingebeni.
Collaborating institutions and agencies
- Africa Centres for Disease Control and Prevention, Addis Ababa, Ethiopia
- Culmen International
- Institute of Ecology and Evolution, University of Edinburgh, UK
- Institute of Tropical Medicine, Antwerp, Belgium
- TransVIHMI, Université de Montpellier, France
- University of Birmingham, UK
- University of California Los-Angeles (UCLA), USA
- Viral Special Pathogens, US Centers for Disease Control and Prevention, USA
- World Health Organization Country Office, Kinshasa, DRC
- World Health Organization, Geneva, Switzerland
References
- Vakaniaki EH, Kacita C, Kinganda-Lusamaki E, et al. Sustained outbreak of a new MPXV lineage in eastern DRC. Nat Med. 2024 Jun 13.
- Wawina-Bokalanga T, Merritt S, Kinganda-Lusamaki E, et al. Epidemiology of distinct mpox outbreaks in Kinshasa: a retrospective study. Lancet. 2025 Jul 5;406(10498):63-75.
- World Health Organization (WHO). WHO response to cholera outbreak in DRC. 2025; Available.
- World Health Organization. Malaria-related respiratory infections in the DRC. 2024. Available.
- Kinganda-Lusamaki, E., et al. 2020 EVD outbreak in Équateur Province, DRC: a genomic characterisation. The Lancet. Microbe, 5(2), e109–e118.
- Katoh K, Rozewicki J, Yamada KD. MAFFT online service for multiple sequence alignment. Briefings in Bioinformatics July 2019.
- Minh BQ, et al. IQ-TREE 2: New methods for phylogenetic inference. Mol Biol Evol. 2020 May 1;37(5):1530-4.